Dissecting the genomic complexity underlying medulloblastoma
- PMID: 22832583
- PMCID: PMC3662966
- DOI: 10.1038/nature11284
Dissecting the genomic complexity underlying medulloblastoma
Abstract
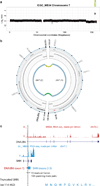
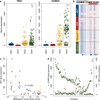
Medulloblastoma is an aggressively growing tumour, arising in the cerebellum or medulla/brain stem. It is the most common malignant brain tumour in children, and shows tremendous biological and clinical heterogeneity. Despite recent treatment advances, approximately 40% of children experience tumour recurrence, and 30% will die from their disease. Those who survive often have a significantly reduced quality of life. Four tumour subgroups with distinct clinical, biological and genetic profiles are currently identified. WNT tumours, showing activated wingless pathway signalling, carry a favourable prognosis under current treatment regimens. SHH tumours show hedgehog pathway activation, and have an intermediate prognosis. Group 3 and 4 tumours are molecularly less well characterized, and also present the greatest clinical challenges. The full repertoire of genetic events driving this distinction, however, remains unclear. Here we describe an integrative deep-sequencing analysis of 125 tumour-normal pairs, conducted as part of the International Cancer Genome Consortium (ICGC) PedBrain Tumor Project. Tetraploidy was identified as a frequent early event in Group 3 and 4 tumours, and a positive correlation between patient age and mutation rate was observed. Several recurrent mutations were identified, both in known medulloblastoma-related genes (CTNNB1, PTCH1, MLL2, SMARCA4) and in genes not previously linked to this tumour (DDX3X, CTDNEP1, KDM6A, TBR1), often in subgroup-specific patterns. RNA sequencing confirmed these alterations, and revealed the expression of what are, to our knowledge, the first medulloblastoma fusion genes identified. Chromatin modifiers were frequently altered across all subgroups. These findings enhance our understanding of the genomic complexity and heterogeneity underlying medulloblastoma, and provide several potential targets for new therapeutics, especially for Group 3 and 4 patients.
Figures


Similar articles
-
Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations.Nature. 2012 Aug 2;488(7409):106-10. doi: 10.1038/nature11329. Nature. 2012. PMID: 22820256 Free PMC article.
-
Novel mutations target distinct subgroups of medulloblastoma.Nature. 2012 Aug 2;488(7409):43-8. doi: 10.1038/nature11213. Nature. 2012. PMID: 22722829 Free PMC article.
-
Medulloblastoma in the age of molecular subgroups: a review.J Neurosurg Pediatr. 2019 Oct 1;24(4):353-363. doi: 10.3171/2019.5.PEDS18381. Epub 2019 Oct 1. J Neurosurg Pediatr. 2019. PMID: 31574483 Review.
-
Rapid diagnosis of medulloblastoma molecular subgroups.Clin Cancer Res. 2011 Apr 1;17(7):1883-94. doi: 10.1158/1078-0432.CCR-10-2210. Epub 2011 Feb 16. Clin Cancer Res. 2011. PMID: 21325292 Free PMC article.
-
Medulloblastoma, WNT-activated/SHH-activated: clinical impact of molecular analysis and histogenetic evaluation.Childs Nerv Syst. 2018 May;34(5):809-815. doi: 10.1007/s00381-018-3765-2. Epub 2018 Mar 26. Childs Nerv Syst. 2018. PMID: 29582169 Review.
Cited by
-
Exome sequencing identifies somatic mutations of DDX3X in natural killer/T-cell lymphoma.Nat Genet. 2015 Sep;47(9):1061-6. doi: 10.1038/ng.3358. Epub 2015 Jul 20. Nat Genet. 2015. PMID: 26192917
-
Transcriptional dysregulation by aberrant enhancer activation and rewiring in cancer.Cancer Sci. 2021 Jun;112(6):2081-2088. doi: 10.1111/cas.14884. Epub 2021 May 1. Cancer Sci. 2021. PMID: 33728716 Free PMC article. Review.
-
Hypo- and Hyper-Assembly Diseases of RNA-Protein Complexes.Trends Mol Med. 2016 Jul;22(7):615-628. doi: 10.1016/j.molmed.2016.05.005. Epub 2016 Jun 3. Trends Mol Med. 2016. PMID: 27263464 Free PMC article. Review.
-
An integrative pan-cancer-wide analysis of epigenetic enzymes reveals universal patterns of epigenomic deregulation in cancer.Genome Biol. 2015 Jul 14;16(1):140. doi: 10.1186/s13059-015-0699-9. Genome Biol. 2015. PMID: 26169266 Free PMC article.
-
PTEN Signaling in the Postnatal Perivascular Progenitor Niche Drives Medulloblastoma Formation.Cancer Res. 2017 Jan 1;77(1):123-133. doi: 10.1158/0008-5472.CAN-16-1991. Epub 2016 Nov 4. Cancer Res. 2017. PMID: 27815386 Free PMC article.
References
-
- Clifford S, et al. Wnt/Wingless Pathway Activation and Chromosome 6 Loss Characterise a Distinct Molecular Sub-Group of Medulloblastomas Associated with a Favourable Prognosis. Cell Cycle. 2006;5:2666–2670. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

