De novo mutations in PLXND1 and REV3L cause Möbius syndrome
- PMID: 26068067
- PMCID: PMC4648025
- DOI: 10.1038/ncomms8199
De novo mutations in PLXND1 and REV3L cause Möbius syndrome
Abstract
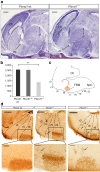
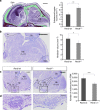
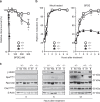
Möbius syndrome (MBS) is a neurological disorder that is characterized by paralysis of the facial nerves and variable other congenital anomalies. The aetiology of this syndrome has been enigmatic since the initial descriptions by von Graefe in 1880 and by Möbius in 1888, and it has been debated for decades whether MBS has a genetic or a non-genetic aetiology. Here, we report de novo mutations affecting two genes, PLXND1 and REV3L in MBS patients. PLXND1 and REV3L represent totally unrelated pathways involved in hindbrain development: neural migration and DNA translesion synthesis, essential for the replication of endogenously damaged DNA, respectively. Interestingly, analysis of Plxnd1 and Rev3l mutant mice shows that disruption of these separate pathways converge at the facial branchiomotor nucleus, affecting either motoneuron migration or proliferation. The finding that PLXND1 and REV3L mutations are responsible for a proportion of MBS patients suggests that de novo mutations in other genes might account for other MBS patients.
Figures

Similar articles
-
Physiopathologic Bases of Moebius Syndrome: Combining Genetic, Vascular, and Teratogenic Theories.Pediatr Neurol. 2024 Apr;153:1-10. doi: 10.1016/j.pediatrneurol.2024.01.007. Epub 2024 Jan 11. Pediatr Neurol. 2024. PMID: 38306744 Review.
-
Developmental delay with hypotrophy associated with homozygous functionally relevant REV3L variant.J Mol Med (Berl). 2021 Mar;99(3):415-423. doi: 10.1007/s00109-020-02033-3. Epub 2021 Jan 20. J Mol Med (Berl). 2021. PMID: 33474647
-
Exploring the Impact of Genetics in a Large Cohort of Moebius Patients by Trio Whole Exome Sequencing.Genes (Basel). 2024 Jul 23;15(8):971. doi: 10.3390/genes15080971. Genes (Basel). 2024. PMID: 39202332 Free PMC article.
-
Multigenic truncation of the semaphorin-plexin pathway by a germline chromothriptic rearrangement associated with Moebius syndrome.Hum Mutat. 2019 Aug;40(8):1057-1062. doi: 10.1002/humu.23775. Epub 2019 May 14. Hum Mutat. 2019. PMID: 31033088 Free PMC article.
-
Poland-Möbius syndrome: a case report implicating a novel mutation of the PLXND1 gene and literature review.BMC Pediatr. 2022 Dec 30;22(1):745. doi: 10.1186/s12887-022-03803-3. BMC Pediatr. 2022. PMID: 36581828 Free PMC article. Review.
Cited by
-
Moebius Syndrome: What We Know So Far.Cureus. 2023 Feb 19;15(2):e35187. doi: 10.7759/cureus.35187. eCollection 2023 Feb. Cureus. 2023. PMID: 36960250 Free PMC article. Review.
-
Characterization of DNA Polymerase Genes in Amazonian Amerindian Populations.Genes (Basel). 2022 Dec 24;14(1):53. doi: 10.3390/genes14010053. Genes (Basel). 2022. PMID: 36672794 Free PMC article.
-
Circulating blood circular RNA in Parkinson's Disease; from involvement in pathology to diagnostic tools in at-risk individuals.NPJ Parkinsons Dis. 2024 Nov 18;10(1):222. doi: 10.1038/s41531-024-00839-3. NPJ Parkinsons Dis. 2024. PMID: 39557914 Free PMC article.
-
MendelVar: gene prioritization at GWAS loci using phenotypic enrichment of Mendelian disease genes.Bioinformatics. 2021 Apr 9;37(1):1-8. doi: 10.1093/bioinformatics/btaa1096. Bioinformatics. 2021. PMID: 33836063 Free PMC article.
-
Möbius syndrome associated with obesity and precocious puberty.BMJ Case Rep. 2018 Dec 7;11(1):e219590. doi: 10.1136/bcr-2017-219590. BMJ Case Rep. 2018. PMID: 30567196 Free PMC article.
References
-
- von Graefe A. In: Handbuch der Gesammten Augenheilkunde eds von Graefe A., Saemisch T. Vol. 6, 148W Engelmann (1880).
-
- Möbius P. J. Ueber angeborene doppelseitige Abducens-Facialis-Lähmung. Münchener Medicinische Wochenschrift. About Congenital 35, 91–94 (1888).
-
- Verzijl H. T., van der Zwaag B., Lammens M., ten Donkelaar H. J. & Padberg G. W. The neuropathology of hereditary congenital facial palsy vs Möbius syndrome. Neurology 64, 649–653 (2005). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

