A mechanism for vertebrate Hedgehog signaling: recruitment to cilia and dissociation of SuFu-Gli protein complexes
- PMID: 20956384
- PMCID: PMC2958481
- DOI: 10.1083/jcb.201004108
A mechanism for vertebrate Hedgehog signaling: recruitment to cilia and dissociation of SuFu-Gli protein complexes
Abstract
In vertebrates, Hedgehog (Hh) signaling initiated in primary cilia activates the membrane protein Smoothened (Smo) and leads to activation of Gli proteins, the transcriptional effectors of the pathway. In the absence of signaling, Gli proteins are inhibited by the cytoplasmic protein Suppressor of Fused (SuFu). It is unclear how Hh activates Gli and whether it directly regulates SuFu. We find that Hh stimulation quickly recruits endogenous SuFu-Gli complexes to cilia, suggesting a model in which Smo activates Gli by relieving inhibition by SuFu. In support of this model, we find that Hh causes rapid dissociation of the SuFu-Gli complex, thus allowing Gli to enter the nucleus and activate transcription. Activation of protein kinase A (PKA), an inhibitor of Hh signaling, blocks ciliary localization of SuFu-Gli complexes, which in turn prevents their dissociation by signaling. Our results support a simple mechanism in which Hh signals at vertebrate cilia cause dissociation of inactive SuFu-Gli complexes, a process inhibited by PKA.
Figures
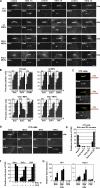
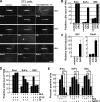
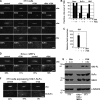

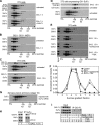

Similar articles
-
Cilium-independent regulation of Gli protein function by Sufu in Hedgehog signaling is evolutionarily conserved.Genes Dev. 2009 Aug 15;23(16):1910-28. doi: 10.1101/gad.1794109. Genes Dev. 2009. PMID: 19684112 Free PMC article.
-
Regulation of Sufu activity by p66β and Mycbp provides new insight into vertebrate Hedgehog signaling.Genes Dev. 2014 Nov 15;28(22):2547-63. doi: 10.1101/gad.249425.114. Genes Dev. 2014. PMID: 25403183 Free PMC article.
-
The ciliary Evc/Evc2 complex interacts with Smo and controls Hedgehog pathway activity in chondrocytes by regulating Sufu/Gli3 dissociation and Gli3 trafficking in primary cilia.Hum Mol Genet. 2013 Jan 1;22(1):124-39. doi: 10.1093/hmg/dds409. Epub 2012 Oct 1. Hum Mol Genet. 2013. PMID: 23026747
-
Hedgehog signaling pathway: a novel model and molecular mechanisms of signal transduction.Cell Mol Life Sci. 2016 Apr;73(7):1317-32. doi: 10.1007/s00018-015-2127-4. Epub 2016 Jan 13. Cell Mol Life Sci. 2016. PMID: 26762301 Free PMC article. Review.
-
Sonic Hedgehog activates the GTPases Rac1 and RhoA in a Gli-independent manner through coupling of smoothened to Gi proteins.Sci Signal. 2011 Nov 22;4(200):pt7. doi: 10.1126/scisignal.2002396. Sci Signal. 2011. PMID: 22114142 Free PMC article. Review.
Cited by
-
The Hedgehog signalling pathway in bone formation.Int J Oral Sci. 2015 Jun 26;7(2):73-9. doi: 10.1038/ijos.2015.14. Int J Oral Sci. 2015. PMID: 26023726 Free PMC article. Review.
-
Members of the Rusc protein family interact with Sufu and inhibit vertebrate Hedgehog signaling.Development. 2016 Nov 1;143(21):3944-3955. doi: 10.1242/dev.138917. Epub 2016 Sep 15. Development. 2016. PMID: 27633991 Free PMC article.
-
Cellular Cholesterol Directly Activates Smoothened in Hedgehog Signaling.Cell. 2016 Aug 25;166(5):1176-1187.e14. doi: 10.1016/j.cell.2016.08.003. Epub 2016 Aug 18. Cell. 2016. PMID: 27545348 Free PMC article.
-
Time-resolved proteomics profiling of the ciliary Hedgehog response.J Cell Biol. 2021 May 3;220(5):e202007207. doi: 10.1083/jcb.202007207. J Cell Biol. 2021. PMID: 33856408 Free PMC article.
-
Bioorthogonal probes for imaging sterols in cells.Chembiochem. 2015 Mar 2;16(4):611-7. doi: 10.1002/cbic.201402715. Epub 2015 Feb 6. Chembiochem. 2015. PMID: 25663046 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

