COG database update 2024
- PMID: 39494517
- PMCID: PMC11701660
- DOI: 10.1093/nar/gkae983
COG database update 2024
Abstract
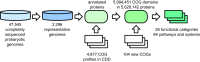
The Clusters of Orthologous Genes (COG) database, originally created in 1997, has been updated to reflect the constantly growing collection of completely sequenced prokaryotic genomes. This update increased the genome coverage from 1309 to 2296 species, including 2103 bacteria and 193 archaea, in most cases, with a single representative genome per genus. This set covers all genera of bacteria and archaea that included organisms with 'complete genomes' as per NCBI databases in November 2023. The number of COGs has been expanded from 4877 to 4981, primarily by including protein families involved in bacterial protein secretion. Accordingly, COG pathways and functional groups now include secretion systems of types II through X, as well as Flp/Tad and type IV pili. These groupings allow straightforward identification and examination of the prokaryotic lineages that encompass-or lack-a particular secretion system. Other developments include improved annotations for the rRNA and tRNA modification proteins, multi-domain signal transduction proteins, and some previously uncharacterized protein families. The new version of COGs is available at https://www.ncbi.nlm.nih.gov/research/COG, as well as on the NCBI FTP site https://ftp.ncbi.nlm.nih.gov/pub/COG/, which also provides archived data from previous COG releases.
Published by Oxford University Press on behalf of Nucleic Acids Research 2024.
Figures
Similar articles
-
Monoclonal Antibody Therapy For High-Risk Coronavirus (COVID 19) Patients With Mild To Moderate Disease Presentations (Archived).2023 Feb 5. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2023 Feb 5. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 34033365 Free Books & Documents.
-
RETRACTED: Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial.Int J Antimicrob Agents. 2020 Jul;56(1):105949. doi: 10.1016/j.ijantimicag.2020.105949. Epub 2020 Mar 20. Int J Antimicrob Agents. 2020. Retraction in: Int J Antimicrob Agents. 2025 Jan;65(1):107416. doi: 10.1016/j.ijantimicag.2024.107416. PMID: 32205204 Free PMC article. Retracted. Clinical Trial.
-
The effectiveness of school-based family asthma educational programs on the quality of life and number of asthma exacerbations of children aged five to 18 years diagnosed with asthma: a systematic review protocol.JBI Database System Rev Implement Rep. 2015 Oct;13(10):69-81. doi: 10.11124/jbisrir-2015-2335. JBI Database System Rev Implement Rep. 2015. PMID: 26571284
-
Depressing time: Waiting, melancholia, and the psychoanalytic practice of care.In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. PMID: 36137063 Free Books & Documents. Review.
-
Conservative, physical and surgical interventions for managing faecal incontinence and constipation in adults with central neurological diseases.Cochrane Database Syst Rev. 2024 Oct 29;10(10):CD002115. doi: 10.1002/14651858.CD002115.pub6. Cochrane Database Syst Rev. 2024. PMID: 39470206
References
-
- Tatusov R.L., Koonin E.V., Lipman D.J.. A genomic perspective on protein families. Science. 1997; 278:631–637. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources