Spectral Algal Fingerprinting and Long Sequencing in Synthetic Algal-Microbial Communities
- PMID: 39329735
- PMCID: PMC11430485
- DOI: 10.3390/cells13181552
Spectral Algal Fingerprinting and Long Sequencing in Synthetic Algal-Microbial Communities
Abstract
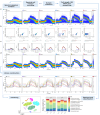
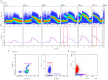
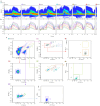
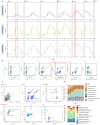
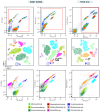
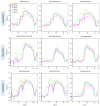
Synthetic biology has advanced in creating artificial microbial and algal communities, but technical and evolutionary complexities still pose significant challenges. Traditional methods, like microscopy and pigment analysis, are limited in throughput and resolution. In contrast, advancements in full-spectrum cytometry enabled high-throughput, multidimensional analysis of single cells based on size, complexity, and spectral fingerprints, offering more precision and flexibility than conventional flow cytometry. This study uses full-spectrum cytometry to analyze synthetic algal-microbial communities, enabling rapid species identification and enumeration. The workflow involves recording individual spectral signatures from monocultures, using autofluorescence to capture populations of interest, and creating a spectral library for further analysis. This spectral library was used for the analysis of the synthetic phytoplankton communities, revealing differences in spectral signatures. Moreover, the synthetic consortium experiment monitored algal growth, comparing results from different instruments, highlighting the advantages of the spectral virtual filter system for precise population separation and abundance tracking. By capturing the entire emission spectrum of each cell, this method enhances understanding of algal-microbial community dynamics and responses to environmental stressors. The development of standardized spectral libraries would improve the characterization of algal communities, further advancing synthetic biology and phytoplankton ecology research.
Keywords: imaging flow cytometry; long sequencing; nanopore-based sequencing; spectral flow cytometry; synthetic algal–microbial communities.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Gallium Scan.2022 Dec 26. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2022 Dec 26. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 33620825 Free Books & Documents.
-
Enabling Systemic Identification and Functionality Profiling for Cdc42 Homeostatic Modulators.bioRxiv [Preprint]. 2024 Jan 8:2024.01.05.574351. doi: 10.1101/2024.01.05.574351. bioRxiv. 2024. Update in: Commun Chem. 2024 Nov 19;7(1):271. doi: 10.1038/s42004-024-01352-7. PMID: 38260445 Free PMC article. Updated. Preprint.
-
Defining the optimum strategy for identifying adults and children with coeliac disease: systematic review and economic modelling.Health Technol Assess. 2022 Oct;26(44):1-310. doi: 10.3310/ZUCE8371. Health Technol Assess. 2022. PMID: 36321689 Free PMC article.
-
Depressing time: Waiting, melancholia, and the psychoanalytic practice of care.In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. PMID: 36137063 Free Books & Documents. Review.
-
Antibody tests for identification of current and past infection with SARS-CoV-2.Cochrane Database Syst Rev. 2022 Nov 17;11(11):CD013652. doi: 10.1002/14651858.CD013652.pub2. Cochrane Database Syst Rev. 2022. PMID: 36394900 Free PMC article. Review.
References
-
- Andersen R.A. Algal Culturing Techniques. Elsevier; Amsterdam, The Netherlands: 2005.
-
- De-Bashan L.E., Bashan Y., Moreno M., Lebsky V.K., Bustillos J.J. Increased Pigment and Lipid Content, Lipid Variety, and Cell and Population Size of the Microalgae Chlorella Spp. When Co-Immobilized in Alginate Beads with the Microalgae-Growth-Promoting Bacterium Azospirillum Brasilense. Can. J. Microbiol. 2002;48:514–521. doi: 10.1139/w02-051. - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

