Molecular characterization of Rft1, an ER membrane protein associated with congenital disorder of glycosylation RFT1-CDG
- PMID: 39025454
- PMCID: PMC11365447
- DOI: 10.1016/j.jbc.2024.107584
Molecular characterization of Rft1, an ER membrane protein associated with congenital disorder of glycosylation RFT1-CDG
Abstract
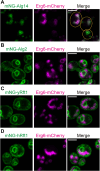
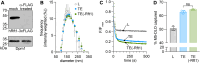
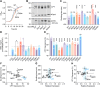
The oligosaccharide needed for protein N-glycosylation is assembled on a lipid carrier via a multistep pathway. Synthesis is initiated on the cytoplasmic face of the endoplasmic reticulum (ER) and completed on the luminal side after transbilayer translocation of a heptasaccharide lipid intermediate. More than 30 congenital disorders of glycosylation (CDGs) are associated with this pathway, including RFT1-CDG which results from defects in the membrane protein Rft1. Rft1 is essential for the viability of yeast and mammalian cells and was proposed as the transporter needed to flip the heptasaccharide lipid intermediate across the ER membrane. However, other studies indicated that Rft1 is not required for heptasaccharide lipid flipping in microsomes or unilamellar vesicles reconstituted with ER membrane proteins, nor is it required for the viability of at least one eukaryote. It is therefore not known what essential role Rft1 plays in N-glycosylation. Here, we present a molecular characterization of human Rft1, using yeast cells as a reporter system. We show that it is a multispanning membrane protein located in the ER, with its N and C termini facing the cytoplasm. It is not N-glycosylated. The majority of RFT1-CDG mutations map to highly conserved regions of the protein. We identify key residues that are important for Rft1's ability to support N-glycosylation and cell viability. Our results provide a necessary platform for future work on this enigmatic protein.
Keywords: CDG; Ist2; MurJ; N-glycosylation; Nvj1; dolichol-linked oligosaccharide; endoplasmic reticulum; lipid droplet; scramblase; yeast.
Copyright © 2024 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures


Update of
-
Molecular characterization of Rft1, an ER membrane protein associated with congenital disorder of glycosylation RFT1-CDG.bioRxiv [Preprint]. 2024 Jun 22:2024.04.03.587922. doi: 10.1101/2024.04.03.587922. bioRxiv. 2024. Update in: J Biol Chem. 2024 Aug;300(8):107584. doi: 10.1016/j.jbc.2024.107584. PMID: 38617304 Free PMC article. Updated. Preprint.
Similar articles
-
Molecular characterization of Rft1, an ER membrane protein associated with congenital disorder of glycosylation RFT1-CDG.bioRxiv [Preprint]. 2024 Jun 22:2024.04.03.587922. doi: 10.1101/2024.04.03.587922. bioRxiv. 2024. Update in: J Biol Chem. 2024 Aug;300(8):107584. doi: 10.1016/j.jbc.2024.107584. PMID: 38617304 Free PMC article. Updated. Preprint.
-
Glycoprotein biosynthesis in a eukaryote lacking the membrane protein Rft1.J Biol Chem. 2013 Jul 12;288(28):20616-23. doi: 10.1074/jbc.M113.479642. Epub 2013 May 28. J Biol Chem. 2013. PMID: 23720757 Free PMC article.
-
RFT1-CDG: deafness as a novel feature of congenital disorders of glycosylation.J Inherit Metab Dis. 2009 Dec;32 Suppl 1:S335-8. doi: 10.1007/s10545-009-1297-3. Epub 2009 Oct 24. J Inherit Metab Dis. 2009. PMID: 19856127
-
Congenital disorders of glycosylation: genetic model systems lead the way.Trends Cell Biol. 2001 Mar;11(3):136-41. doi: 10.1016/s0962-8924(01)01925-0. Trends Cell Biol. 2001. PMID: 11306275 Review.
-
SSR4-CDG, an ultra-rare X-linked congenital disorder of glycosylation affecting the TRAP complex: Review of 22 affected individuals including the first adult patient.Mol Genet Metab. 2024 Jul;142(3):108477. doi: 10.1016/j.ymgme.2024.108477. Epub 2024 Apr 18. Mol Genet Metab. 2024. PMID: 38805916 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

