Insight into the mechanism of gallstone disease by proteomic and metaproteomic characterization of human bile
- PMID: 38111640
- PMCID: PMC10726133
- DOI: 10.3389/fmicb.2023.1276951
Insight into the mechanism of gallstone disease by proteomic and metaproteomic characterization of human bile
Abstract
Introduction: Cholesterol gallstone disease is a prevalent condition that has a significant economic impact. However, the role of the bile microbiome in its development and the host's responses to it remain poorly understood.
Methods: In this study, we conducted a comprehensive analysis of microbial and human bile proteins in 40 individuals with either gallstone disease or gallbladder polyps. We employed a combined proteomic and metaproteomic approach, as well as meta-taxonomic analysis, functional pathway enrichment, and Western blot analyses.
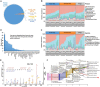
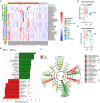
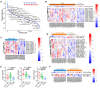
Results: Our metaproteomic analysis, utilizing the lowest common ancestor algorithm, identified 158 microbial taxa in the bile samples. We discovered microbial taxa that may contribute to gallstone formation, including β-glucuronidase-producing bacteria such as Streptococcus, Staphylococcus, and Clostridium, as well as those involved in biofilm formation like Helicobacter, Cyanobacteria, Pseudomonas, Escherichia coli, and Clostridium. Furthermore, we identified 2,749 human proteins and 87 microbial proteins with a protein false discovery rate (FDR) of 1% and at least 2 distinct peptides. Among these proteins, we found microbial proteins crucial to biofilm formation, such as QDR3, ompA, ndk, pstS, nanA, pfIB, and dnaK. Notably, QDR3 showed a gradual upregulation from chronic to acute cholesterol gallstone disease when compared to polyp samples. Additionally, we discovered other microbial proteins that enhance bacterial virulence and gallstone formation by counteracting host oxidative stress, including sodB, katG, rbr, htrA, and ahpC. We also identified microbial proteins like lepA, rtxA, pckA, tuf, and tpiA that are linked to bacterial virulence and potential gallstone formation, with lepA being upregulated in gallstone bile compared to polyp bile. Furthermore, our analysis of the host proteome in gallstone bile revealed enhanced inflammatory molecular profiles, including innate immune molecules against microbial infections. Gallstone bile exhibited overrepresented pathways related to blood coagulation, folate metabolism, and the IL-17 pathway. However, we observed suppressed metabolic activities, particularly catabolic metabolism and transport activities, in gallstone bile compared to polyp bile. Notably, acute cholelithiasis bile demonstrated significantly impaired metabolic activities compared to chronic cholelithiasis bile.
Conclusion: Our study provides a comprehensive metaproteomic analysis of bile samples related to gallstone disease, offering new insights into the microbiome-host interaction and gallstone formation mechanism.
Keywords: bile; cholecystitis; gallstone; metaproteomics; microbiome; proteomics.
Copyright © 2023 Yang, Wang, Jiang, Zhang, Du, Li and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Landscape in the gallbladder mycobiome and bacteriome of patients undergoing cholelithiasis with chronic cholecystitis.Front Microbiol. 2023 Mar 22;14:1131694. doi: 10.3389/fmicb.2023.1131694. eCollection 2023. Front Microbiol. 2023. PMID: 37032855 Free PMC article.
-
Increasing taxonomic and functional characterization of host-microbiome interactions by DIA-PASEF metaproteomics.Front Microbiol. 2023 Oct 16;14:1258703. doi: 10.3389/fmicb.2023.1258703. eCollection 2023. Front Microbiol. 2023. PMID: 37908546 Free PMC article.
-
Metagenomic sequencing of bile from gallstone patients to identify different microbial community patterns and novel biliary bacteria.Sci Rep. 2015 Dec 2;5:17450. doi: 10.1038/srep17450. Sci Rep. 2015. PMID: 26625708 Free PMC article.
-
Role of the biliary microbiome in gallstone disease.Expert Rev Gastroenterol Hepatol. 2018 Dec;12(12):1193-1205. doi: 10.1080/17474124.2018.1533812. Epub 2018 Oct 14. Expert Rev Gastroenterol Hepatol. 2018. PMID: 30791792 Review.
-
Triglycerides and gallstone formation.Clin Chim Acta. 2010 Nov 11;411(21-22):1625-31. doi: 10.1016/j.cca.2010.08.003. Epub 2010 Aug 10. Clin Chim Acta. 2010. PMID: 20699090 Review.
Cited by
-
Proteomics and metabolomics analyses of urine for investigation of gallstone disease in a high-altitude area.Metabolomics. 2024 Aug 14;20(5):99. doi: 10.1007/s11306-024-02162-7. Metabolomics. 2024. PMID: 39143352
-
The role of bacteria in gallstone formation.Folia Microbiol (Praha). 2024 Feb;69(1):33-40. doi: 10.1007/s12223-024-01131-w. Epub 2024 Jan 22. Folia Microbiol (Praha). 2024. PMID: 38252338 Review.
References
-
- Barahona Ponce C., Scherer D., Brinster R., Boekstegers F., Marcelain K., Garate-Calderon V., et al. . (2021). Gallstones, body mass index, C-reactive protein, and gallbladder Cancer: Mendelian randomization analysis of Chilean and European genotype data. Hepatology 73, 1783–1796. doi: 10.1002/hep.31537, PMID: - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

