Bile Microbiome Signatures Associated with Pancreatic Ductal Adenocarcinoma Compared to Benign Disease: A UK Pilot Study
- PMID: 38069211
- PMCID: PMC10706407
- DOI: 10.3390/ijms242316888
Bile Microbiome Signatures Associated with Pancreatic Ductal Adenocarcinoma Compared to Benign Disease: A UK Pilot Study
Abstract
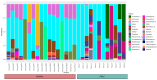
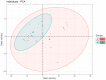
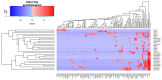
Pancreatic ductal adenocarcinoma (PDAC) has a very poor survival. The intra-tumoural microbiome can influence pancreatic tumourigenesis and chemoresistance and, therefore, patient survival. The role played by bile microbiota in PDAC is unknown. We aimed to define bile microbiome signatures that can effectively distinguish malignant from benign tumours in patients presenting with obstructive jaundice caused by benign and malignant pancreaticobiliary disease. Prospective bile samples were obtained from 31 patients who underwent either Endoscopic Retrograde Cholangiopancreatography (ERCP) or Percutaneous Transhepatic Cholangiogram (PTC). Variable regions (V3-V4) of the 16S rRNA genes of microorganisms present in the samples were amplified by Polymerase Chain Reaction (PCR) and sequenced. The cohort consisted of 12 PDAC, 10 choledocholithiasis, seven gallstone pancreatitis and two primary sclerosing cholangitis patients. Using the 16S rRNA method, we identified a total of 135 genera from 29 individuals (12 PDAC and 17 benign). The bile microbial beta diversity significantly differed between patients with PDAC vs. benign disease (Permanova p = 0.0173). The separation of PDAC from benign samples is clearly seen through unsupervised clustering of Aitchison distance. We found three genera to be of significantly lower abundance among PDAC samples vs. benign, adjusting for false discovery rate (FDR). These were Escherichia (FDR = 0.002) and two unclassified genera, one from Proteobacteria (FDR = 0.002) and one from Enterobacteriaceae (FDR = 0.011). In the same samples, the genus Streptococcus (FDR = 0.033) was found to be of increased abundance in the PDAC group. We show that patients with obstructive jaundice caused by PDAC have an altered microbiome composition in the bile compared to those with benign disease. These bile-based microbes could be developed into potential diagnostic and prognostic biomarkers for PDAC and warrant further investigation.
Keywords: 16S rRNA gene; bile; biomarker; microbiome; pancreatic cancer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Alterations of the bile microbiome is associated with progression-free survival in pancreatic ductal adenocarcinoma patients.BMC Microbiol. 2024 Jul 1;24(1):235. doi: 10.1186/s12866-024-03371-9. BMC Microbiol. 2024. PMID: 38956452 Free PMC article.
-
Endoscopic ultrasound-guided fine-needle biopsy as a tool for studying the intra-tumoral microbiome in pancreatic ductal adenocarcinoma: a pilot study.Sci Rep. 2022 Jan 7;12(1):107. doi: 10.1038/s41598-021-04095-w. Sci Rep. 2022. PMID: 34997106 Free PMC article.
-
Microbiomic profiles of bile in patients with benign and malignant pancreaticobiliary disease.PLoS One. 2023 Apr 18;18(4):e0283021. doi: 10.1371/journal.pone.0283021. eCollection 2023. PLoS One. 2023. PMID: 37071646 Free PMC article.
-
MICRObiota on BILIOpancreatic malignant diseases [MICROBILIO]: A systematic review.Clinics (Sao Paulo). 2022 Sep 16;77:100101. doi: 10.1016/j.clinsp.2022.100101. eCollection 2022. Clinics (Sao Paulo). 2022. PMID: 36122499 Free PMC article. Review.
-
Role of the microbiome in systemic therapy for pancreatic ductal adenocarcinoma (Review).Int J Oncol. 2021 Dec;59(6):101. doi: 10.3892/ijo.2021.5281. Epub 2021 Nov 5. Int J Oncol. 2021. PMID: 34738624 Free PMC article. Review.
Cited by
-
Emerging Therapeutic Options in Pancreatic Cancer Management.Int J Mol Sci. 2024 Feb 5;25(3):1929. doi: 10.3390/ijms25031929. Int J Mol Sci. 2024. PMID: 38339207 Free PMC article.
-
MicroRNAs as Bile-based biomarkers in pancreaticobiliary cancers (MIRABILE): a cohort study.Int J Surg. 2024 Oct 1;110(10):6518-6527. doi: 10.1097/JS9.0000000000001888. Int J Surg. 2024. PMID: 39041944 Free PMC article.
-
The microbial composition of pancreatic ductal adenocarcinoma: a systematic review of 16S rRNA gene sequencing.Int J Surg. 2024 Oct 1;110(10):6771-6799. doi: 10.1097/JS9.0000000000001762. Int J Surg. 2024. PMID: 38874485 Free PMC article.
-
Intratumoral Microbiota: Metabolic Influences and Biomarker Potential in Gastrointestinal Cancer.Biomolecules. 2024 Jul 27;14(8):917. doi: 10.3390/biom14080917. Biomolecules. 2024. PMID: 39199305 Free PMC article. Review.
References
-
- Neoptolemos J.P., Palmer D.H., Ghaneh P., Psarelli E.E., Valle J.W., Halloran C.M., Faluyi O., O’Reilly D.A., Cunningham D., Wadsley J., et al. Comparison of adjuvant gemcitabine and capecitabine with gemcitabine monotherapy in patients with resected pancreatic cancer (ESPAC-4): A multicentre, open-label, randomised, phase 3 trial. Lancet. 2017;389:1011–1024. doi: 10.1016/S0140-6736(16)32409-6. - DOI - PubMed
-
- Pushalkar S., Hundeyin M., Daley D., Zambirinis C.P., Kurz E., Mishra A., Mohan N., Aykut B., Usyk M., Torres L.E., et al. The Pancreatic Cancer Microbiome Promotes Oncogenesis by Induction of Innate and Adaptive Immune Suppression. Cancer Discov. 2018;8:403–416. doi: 10.1158/2159-8290.CD-17-1134. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

