Validation of lipid-related therapeutic targets for coronary heart disease prevention using human genetics
- PMID: 34675202
- PMCID: PMC8531035
- DOI: 10.1038/s41467-021-25731-z
Validation of lipid-related therapeutic targets for coronary heart disease prevention using human genetics
Abstract
Drug target Mendelian randomization (MR) studies use DNA sequence variants in or near a gene encoding a drug target, that alter the target's expression or function, as a tool to anticipate the effect of drug action on the same target. Here we apply MR to prioritize drug targets for their causal relevance for coronary heart disease (CHD). The targets are further prioritized using independent replication, co-localization, protein expression profiles and data from the British National Formulary and clinicaltrials.gov. Out of the 341 drug targets identified through their association with blood lipids (HDL-C, LDL-C and triglycerides), we robustly prioritize 30 targets that might elicit beneficial effects in the prevention or treatment of CHD, including NPC1L1 and PCSK9, the targets of drugs used in CHD prevention. We discuss how this approach can be generalized to other targets, disease biomarkers and endpoints to help prioritize and validate targets during the drug development process.
© 2021. The Author(s).
Conflict of interest statement
A.F.S. has received Servier funding for unrelated work. M.Z. conducted this research as an employee of BenevolentAI. Since completing the work M.Z. is now a full-time employee of GlaxoSmithKline.
Figures

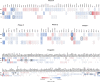
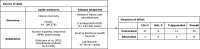
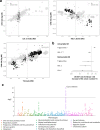
Similar articles
-
Effects of genetically proxied lipid-lowering drugs on acute myocardial infarction: a drug-target mendelian randomization study.Lipids Health Dis. 2024 Jun 3;23(1):163. doi: 10.1186/s12944-024-02133-w. Lipids Health Dis. 2024. PMID: 38831433 Free PMC article.
-
Genetic association of lipids and lipid lowering drug target genes with sarcoidosis.Sci Rep. 2024 Oct 17;14(1):24351. doi: 10.1038/s41598-024-75322-3. Sci Rep. 2024. PMID: 39420052 Free PMC article.
-
Association of genetically predicted lipid traits and lipid-modifying targets with heart failure.Eur J Prev Cardiol. 2023 Mar 1;30(4):358-366. doi: 10.1093/eurjpc/zwac290. Eur J Prev Cardiol. 2023. PMID: 36520639
-
Mendelian randomization to assess causal effects of blood lipids on coronary heart disease: lessons from the past and applications to the future.Curr Opin Endocrinol Diabetes Obes. 2016 Apr;23(2):124-30. doi: 10.1097/MED.0000000000000230. Curr Opin Endocrinol Diabetes Obes. 2016. PMID: 26910273 Free PMC article. Review.
-
Pharmacogenomics of blood lipid regulation.Pharmacogenomics. 2018 May;19(7):651-665. doi: 10.2217/pgs-2018-0007. Epub 2018 Apr 30. Pharmacogenomics. 2018. PMID: 29706123 Review.
Cited by
-
Network pharmacology combined with Mendelian randomization analysis to identify the key targets of renin-angiotensin-aldosterone system inhibitors in the treatment of diabetic nephropathy.Front Endocrinol (Lausanne). 2024 Jan 25;15:1354950. doi: 10.3389/fendo.2024.1354950. eCollection 2024. Front Endocrinol (Lausanne). 2024. PMID: 38332893 Free PMC article.
-
Effect of genetically predicted sclerostin on cardiovascular biomarkers, risk factors, and disease outcomes.Nat Commun. 2024 Nov 13;15(1):9832. doi: 10.1038/s41467-024-53623-5. Nat Commun. 2024. PMID: 39537602 Free PMC article.
-
Mendelian randomization studies on coronary artery disease: a systematic review and meta-analysis.Syst Rev. 2024 Jan 16;13(1):29. doi: 10.1186/s13643-023-02442-8. Syst Rev. 2024. PMID: 38225600 Free PMC article.
-
Systematic Mendelian Randomization Exploring Druggable Genes for Hemorrhagic Strokes.Mol Neurobiol. 2025 Feb;62(2):1359-1372. doi: 10.1007/s12035-024-04336-9. Epub 2024 Jul 9. Mol Neurobiol. 2025. PMID: 38977622 Free PMC article.
-
Multivariate genome-wide analysis of aging-related traits identifies novel loci and new drug targets for healthy aging.Nat Aging. 2023 Aug;3(8):1020-1035. doi: 10.1038/s43587-023-00455-5. Epub 2023 Aug 7. Nat Aging. 2023. PMID: 37550455 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- RG/10/12/28456/BHF_/British Heart Foundation/United Kingdom
- CH/F/20/90003/BHF_/British Heart Foundation/United Kingdom
- FS/17/70/33482/BHF_/British Heart Foundation/United Kingdom
- MR/R024227/1/MRC_/Medical Research Council/United Kingdom
- S011676/MRC_/Medical Research Council/United Kingdom
- PG/18/50/33837/BHF_/British Heart Foundation/United Kingdom
- MR/V033867/1/MRC_/Medical Research Council/United Kingdom
- R01 AG056477/AG/NIA NIH HHS/United States
- MC_UU_00011/6/MRC_/Medical Research Council/United Kingdom
- MC_UU_00011/4/MRC_/Medical Research Council/United Kingdom
- RG/19/4/34452/BHF_/British Heart Foundation/United Kingdom
- 221854/Z/20/Z/WT_/Wellcome Trust/United Kingdom
- 29019/CRUK_/Cancer Research UK/United Kingdom
- RF1 AG062553/AG/NIA NIH HHS/United States
- SP/13/6/30554/BHF_/British Heart Foundation/United Kingdom
- MR/S011676/1/MRC_/Medical Research Council/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- R024227/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Miscellaneous

