Cooperation, Competition, and Specialized Metabolism in a Simplified Root Nodule Microbiome
- PMID: 32843548
- PMCID: PMC7448283
- DOI: 10.1128/mBio.01917-20
Cooperation, Competition, and Specialized Metabolism in a Simplified Root Nodule Microbiome
Abstract
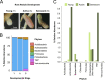
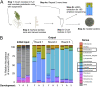
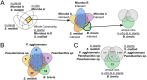
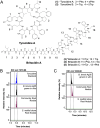
Microbiomes associated with various plant structures often contain members with the potential to make specialized metabolites, e.g., molecules with antibacterial, antifungal, or siderophore activities. However, when and where microbes associated with plants produce specialized metabolites, and the potential role of these molecules in mediating intramicrobiome interactions, is not well understood. Root nodules of legume plants are organs devoted to hosting symbiotic bacteria that fix atmospheric nitrogen and have recently been shown to harbor a relatively simple accessory microbiome containing members with the ability to produce specialized metabolites in vitro On the basis of these observations, we sought to develop a model nodule microbiome system for evaluating specialized microbial metabolism in planta Starting with an inoculum derived from field-grown Medicago sativa nodules, serial passaging through gnotobiotic nodules yielded a simplified accessory community composed of four members: Brevibacillus brevis, Paenibacillus sp., Pantoea agglomerans, and Pseudomonas sp. Some members of this community exhibited clear cooperation in planta, while others were antagonistic and capable of disrupting cooperation between other partners. Using matrix-assisted laser desorption ionization-imaging mass spectrometry, we found that metabolites associated with individual taxa had unique distributions, indicating that some members of the nodule community were spatially segregated. Finally, we identified two families of molecules produced by B. brevisin planta as the antibacterial tyrocidines and a novel set of gramicidin-type molecules, which we term the britacidins. Collectively, these results indicate that in addition to nitrogen fixation, legume root nodules are likely also sites of active antimicrobial production.
Keywords: antibiotic; microbial interactions; microbiome; root nodule; specialized metabolism.
Copyright © 2020 Hansen et al.
Figures



Similar articles
-
Characterization of the Sinorhizobium meliloti HslUV and ClpXP Protease Systems in Free-Living and Symbiotic States.J Bacteriol. 2019 Mar 13;201(7):e00498-18. doi: 10.1128/JB.00498-18. Print 2019 Apr 1. J Bacteriol. 2019. PMID: 30670545 Free PMC article.
-
MALDI mass spectrometry-assisted molecular imaging of metabolites during nitrogen fixation in the Medicago truncatula-Sinorhizobium meliloti symbiosis.Plant J. 2013 Jul;75(1):130-145. doi: 10.1111/tpj.12191. Epub 2013 May 6. Plant J. 2013. PMID: 23551619
-
Two cultivated legume plants reveal the enrichment process of the microbiome in the rhizocompartments.Mol Ecol. 2017 Mar;26(6):1641-1651. doi: 10.1111/mec.14027. Epub 2017 Feb 23. Mol Ecol. 2017. PMID: 28139080
-
Mechanisms underlying legume-rhizobium symbioses.J Integr Plant Biol. 2022 Feb;64(2):244-267. doi: 10.1111/jipb.13207. J Integr Plant Biol. 2022. PMID: 34962095 Review.
-
Gene Expression in Nitrogen-Fixing Symbiotic Nodule Cells in Medicago truncatula and Other Nodulating Plants.Plant Cell. 2020 Jan;32(1):42-68. doi: 10.1105/tpc.19.00494. Epub 2019 Nov 11. Plant Cell. 2020. PMID: 31712407 Free PMC article. Review.
Cited by
-
Nodule-associated diazotrophic community succession is driven by developmental phases combined with microhabitat of Sophora davidii.Front Microbiol. 2022 Dec 1;13:1078208. doi: 10.3389/fmicb.2022.1078208. eCollection 2022. Front Microbiol. 2022. PMID: 36532429 Free PMC article.
-
Both incompatible and compatible rhizobia inhabit the intercellular spaces of leguminous root nodules.Plant Signal Behav. 2023 Dec 31;18(1):2245995. doi: 10.1080/15592324.2023.2245995. Plant Signal Behav. 2023. PMID: 37573516 Free PMC article.
-
Competition, Nodule Occupancy, and Persistence of Inoculant Strains: Key Factors in the Rhizobium-Legume Symbioses.Front Plant Sci. 2021 Aug 19;12:690567. doi: 10.3389/fpls.2021.690567. eCollection 2021. Front Plant Sci. 2021. PMID: 34489993 Free PMC article. Review.
-
Investigation of the potential of Brevibacillus spp. for the biosynthesis of nonribosomally produced bioactive compounds by combination of genome mining with MALDI-TOF mass spectrometry.Front Microbiol. 2023 Dec 14;14:1286565. doi: 10.3389/fmicb.2023.1286565. eCollection 2023. Front Microbiol. 2023. PMID: 38156002 Free PMC article.
-
Community standards and future opportunities for synthetic communities in plant-microbiota research.Nat Microbiol. 2024 Nov;9(11):2774-2784. doi: 10.1038/s41564-024-01833-4. Epub 2024 Oct 30. Nat Microbiol. 2024. PMID: 39478084 Review.
References
-
- Bacon CW, White JF. 2016. Functions, mechanisms and regulation of endophytic and epiphytic microbial communities of plants. Symbiosis 68:87–98. doi:10.1007/s13199-015-0350-2. - DOI

