Ultrasensitive Response of Developing Myxococcus xanthus to the Addition of Nutrient Medium Correlates with the Level of MrpC
- PMID: 30181127
- PMCID: PMC6199472
- DOI: 10.1128/JB.00456-18
Ultrasensitive Response of Developing Myxococcus xanthus to the Addition of Nutrient Medium Correlates with the Level of MrpC
Abstract
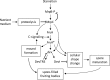
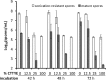
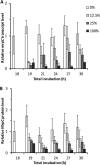
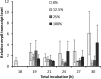
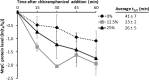
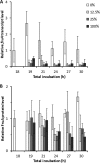
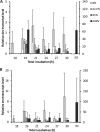
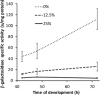
Upon depletion of nutrients, Myxococcus xanthus forms mounds on a solid surface. The differentiation of rod-shaped cells into stress-resistant spores within mounds creates mature fruiting bodies. The developmental process can be perturbed by the addition of nutrient medium before the critical period of commitment to spore formation. The response was investigated by adding a 2-fold dilution series of nutrient medium to starving cells. An ultrasensitive response was observed, as indicated by a steep increase in the spore number after the addition of 12.5% versus 25% nutrient medium. The level of MrpC, which is a key transcription factor in the gene regulatory network, correlated with the spore number after nutrient medium addition. The MrpC level decreased markedly by 3 h after adding nutrient medium but recovered more after the addition of 12.5% than after 25% nutrient medium addition. The difference in MrpC levels was greatest midway during the period of commitment to sporulation, and mound formation was restored after 12.5% nutrient medium addition but not after adding 25% nutrient medium. Although the number of spores formed after 12.5% nutrient medium addition was almost normal, the transcript levels of "late" genes in the regulatory network failed to rise normally during the commitment period. However, at later times, expression from a reporter gene fused to a late promoter was higher after adding 12.5% than after adding 25% nutrient medium, consistent with the spore numbers. The results suggest that a threshold level of MrpC must be achieved in order for mounds to persist and for cells within to differentiate into spores.IMPORTANCE Many signaling and gene regulatory networks convert graded stimuli into all-or-none switch-like responses. Such ultrasensitivity can produce bistability in cell populations, leading to different cell fates and enhancing survival. We discovered an ultrasensitive response of M. xanthus to nutrient medium addition during development. A small change in nutrient medium concentration caused a profound change in the developmental process. The level of the transcription factor MrpC correlated with multicellular mound formation and differentiation into spores. A threshold level of MrpC is proposed to be necessary to initiate mound formation and create a positive feedback loop that may explain the ultrasensitive response. Understanding how this biological switch operates will provide a paradigm for the broadly important topic of cellular behavior in microbial communities.
Keywords: FruA; MrpC; Myxococcus xanthus; bacterial development; gene regulatory network; sporulation; ultrasensitive response.
Copyright © 2018 American Society for Microbiology.
Figures

Similar articles
-
Nutrient-regulated proteolysis of MrpC halts expression of genes important for commitment to sporulation during Myxococcus xanthus development.J Bacteriol. 2014 Aug;196(15):2736-47. doi: 10.1128/JB.01692-14. Epub 2014 May 16. J Bacteriol. 2014. PMID: 24837289 Free PMC article.
-
Transcription factor MrpC binds to promoter regions of hundreds of developmentally-regulated genes in Myxococcus xanthus.BMC Genomics. 2014 Dec 16;15:1123. doi: 10.1186/1471-2164-15-1123. BMC Genomics. 2014. PMID: 25515642 Free PMC article.
-
Systematic analysis of the Myxococcus xanthus developmental gene regulatory network supports posttranslational regulation of FruA by C-signaling.Mol Microbiol. 2019 Jun;111(6):1732-1752. doi: 10.1111/mmi.14249. Epub 2019 Apr 6. Mol Microbiol. 2019. PMID: 30895656
-
Dual regulation with Ser/Thr kinase cascade and a His/Asp TCS in Myxococcus xanthus.Adv Exp Med Biol. 2008;631:111-21. doi: 10.1007/978-0-387-78885-2_7. Adv Exp Med Biol. 2008. PMID: 18792684 Review.
-
Two-Component Signal Transduction Systems That Regulate the Temporal and Spatial Expression of Myxococcus xanthus Sporulation Genes.J Bacteriol. 2015 Sep 14;198(3):377-85. doi: 10.1128/JB.00474-15. Print 2016 Feb 1. J Bacteriol. 2015. PMID: 26369581 Free PMC article. Review.
Cited by
-
Transcriptomic analysis of the Myxococcus xanthus FruA regulon, and comparative developmental transcriptomic analysis of two fruiting body forming species, Myxococcus xanthus and Myxococcus stipitatus.BMC Genomics. 2021 Nov 1;22(1):784. doi: 10.1186/s12864-021-08051-w. BMC Genomics. 2021. PMID: 34724903 Free PMC article.
-
Mutation of self-binding sites in the promoter of the MrpC transcriptional regulator leads to asynchronous Myxococcus xanthus development.Front Microbiol. 2023 Nov 23;14:1293966. doi: 10.3389/fmicb.2023.1293966. eCollection 2023. Front Microbiol. 2023. PMID: 38075919 Free PMC article.
References
-
- Yang Z, Higgs P. 2014. Myxobacteria: genomics, cellular and molecular biology. Caister Academic Press, Norfolk, UK.
-
- Lee B, Holkenbrink C, Treuner-Lange A, Higgs PI. 2012. Myxococcus xanthus developmental cell fate production: heterogeneous accumulation of developmental regulatory proteins and reexamination of the role of MazF in developmental lysis. J Bacteriol 194:3058–3068. doi:10.1128/JB.06756-11. - DOI - PMC - PubMed
-
- Saha S, Patra P, Igoshin OA, Kroos L. 2018. Systematic analysis of the Myxococcus xanthus developmental gene regulatory network supports posttranslational regulation of FruA by C-signaling. bioRxiv http://biorxiv.org/cgi/content/short/415331v1. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

