Convergence between the microcosms of Southeast Asian and North American pitcher plants
- PMID: 30152327
- PMCID: PMC6130972
- DOI: 10.7554/eLife.36741
Convergence between the microcosms of Southeast Asian and North American pitcher plants
Abstract
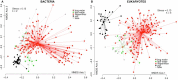
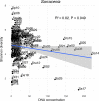
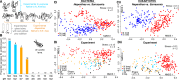
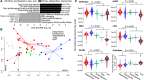
The 'pitchers' of carnivorous pitcher plants are exquisite examples of convergent evolution. An open question is whether the living communities housed in pitchers also converge in structure or function. Using samples from more than 330 field-collected pitchers of eight species of Southeast Asian Nepenthes and six species of North American Sarracenia, we demonstrate that the pitcher microcosms, or miniature ecosystems with complex communities, are strikingly similar. Compared to communities from surrounding habitats, pitcher communities house fewer species. While communities associated with the two genera contain different microbial organisms and arthropods, the species are predominantly from the same phylogenetic clades. Microbiomes from both genera are enriched in degradation pathways and have high abundances of key degradation enzymes. Moreover, in a manipulative field experiment, Nepenthes pitchers placed in a North American bog assembled Sarracenia-like communities. An understanding of the convergent interactions in pitcher microcosms facilitates identification of selective pressures shaping the communities.
Keywords: Nepenthes; Sarracenia; communities; ecology; evolutionary biology; microbial ecology; species interactions.
© 2018, Bittleston et al.
Conflict of interest statement
LB, CW, BY, XC, KC, NP, AP No competing interests declared
Figures





Similar articles
-
Selective Bacterial Community Enrichment between the Pitcher Plants Sarracenia minor and Sarracenia flava.Microbiol Spectr. 2021 Dec 22;9(3):e0069621. doi: 10.1128/Spectrum.00696-21. Epub 2021 Nov 24. Microbiol Spectr. 2021. PMID: 34817222 Free PMC article.
-
Carnivorous Nepenthes Pitchers with Less Acidic Fluid House Nitrogen-Fixing Bacteria.Appl Environ Microbiol. 2023 Jul 26;89(7):e0081223. doi: 10.1128/aem.00812-23. Epub 2023 Jun 20. Appl Environ Microbiol. 2023. PMID: 37338413 Free PMC article.
-
Characterization and Comparison of Convergence Among Cephalotus follicularis Pitcher Plant-Associated Communities With Those of Nepenthes and Sarracenia Found Worldwide.Front Plant Sci. 2022 Jun 6;13:887635. doi: 10.3389/fpls.2022.887635. eCollection 2022. Front Plant Sci. 2022. PMID: 35734258 Free PMC article.
-
Traps of carnivorous pitcher plants as a habitat: composition of the fluid, biodiversity and mutualistic activities.Ann Bot. 2011 Feb;107(2):181-94. doi: 10.1093/aob/mcq238. Epub 2010 Dec 15. Ann Bot. 2011. PMID: 21159782 Free PMC article. Review.
-
Convergent and divergent evolution in carnivorous pitcher plant traps.New Phytol. 2018 Feb;217(3):1035-1041. doi: 10.1111/nph.14879. Epub 2017 Nov 13. New Phytol. 2018. PMID: 29131340 Review.
Cited by
-
On the move: sloths and their epibionts as model mobile ecosystems.Biol Rev Camb Philos Soc. 2021 Dec;96(6):2638-2660. doi: 10.1111/brv.12773. Epub 2021 Jul 26. Biol Rev Camb Philos Soc. 2021. PMID: 34309191 Free PMC article. Review.
-
Convergent evolution in the genomics era: new insights and directions.Philos Trans R Soc Lond B Biol Sci. 2019 Jul 22;374(1777):20190102. doi: 10.1098/rstb.2019.0102. Epub 2019 Jun 3. Philos Trans R Soc Lond B Biol Sci. 2019. PMID: 31154976 Free PMC article. No abstract available.
-
An acidophilic fungus promotes prey digestion in a carnivorous plant.Nat Microbiol. 2024 Oct;9(10):2522-2537. doi: 10.1038/s41564-024-01766-y. Epub 2024 Aug 1. Nat Microbiol. 2024. PMID: 39090391 Free PMC article.
-
Tropical pitcher plants (Nepenthes) act as ecological filters by altering properties of their fluid microenvironments.Sci Rep. 2020 Mar 10;10(1):4431. doi: 10.1038/s41598-020-61193-x. Sci Rep. 2020. PMID: 32157122 Free PMC article.
-
Acid or base? How do plants regulate the ecology of their phylloplane?AoB Plants. 2021 Jun 10;13(4):plab032. doi: 10.1093/aobpla/plab032. eCollection 2021 Aug. AoB Plants. 2021. PMID: 34285793 Free PMC article. Review.
References
-
- Abubucker S, Segata N, Goll J, Schubert AM, Izard J, Cantarel BL, Rodriguez-Mueller B, Zucker J, Thiagarajan M, Henrissat B, White O, Kelley ST, Methé B, Schloss PD, Gevers D, Mitreva M, Huttenhower C. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Computational Biology. 2012;8:e1002358. doi: 10.1371/journal.pcbi.1002358. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- NSF Graduate Fellowship and Doctoral Dissertation Improvement Grant DEB-1400982/National Science Foundation/International
- Foundational Questions In Evolutionary Biology/John Templeton Foundation/International
- Putnam Expedition Grant/Harvard University Museum of Comparative Zoology/International
- High Impact Research Grant, UM-MOHE HIR Grant UM.C/625/1/HIR/MOHE/CHAN/14/1/Universiti Malaya/International
- SES-0750480/National Science Foundation/International
- High Impact Research Grant, H-50001-A000027/Universiti Malaya/International
- High Impact Research Grant, A-000001-50001/Universiti Malaya/International
- Foundational Questions In Evolutionary Biology/Templeton Foundation/International
- Putnam Grant/Harvard University Museum of Comparative Zoology Putnam Expedition Grant/International
- UM-MOHE HIR Grant UM.C/625/1/HIR/MOHE/CHAN/14/1/University of Malaya High Impact Research Grant/International
- H-50001-A000027/University of Malaya High Impact Research Grant/International
- A-000001-50001/University of Malaya High Impact Research Grant/International
LinkOut - more resources
Full Text Sources
Other Literature Sources

