Structural insights into POT1-TPP1 interaction and POT1 C-terminal mutations in human cancer
- PMID: 28393832
- PMCID: PMC5394241
- DOI: 10.1038/ncomms14929
Structural insights into POT1-TPP1 interaction and POT1 C-terminal mutations in human cancer
Abstract
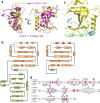
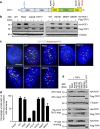
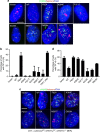
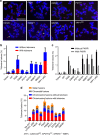
Mammalian shelterin proteins POT1 and TPP1 form a stable heterodimer that protects chromosome ends and regulates telomerase-mediated telomere extension. However, how POT1 interacts with TPP1 remains unknown. Here we present the crystal structure of the C-terminal portion of human POT1 (POT1C) complexed with the POT1-binding motif of TPP1. The structure shows that POT1C contains two domains, a third OB fold and a Holliday junction resolvase-like domain. Both domains are essential for binding to TPP1. Notably, unlike the heart-shaped structure of ciliated protozoan Oxytricha nova TEBPα-β complex, POT1-TPP1 adopts an elongated V-shaped conformation. In addition, we identify several missense mutations in human cancers that disrupt the POT1C-TPP1 interaction, resulting in POT1 instability. POT1C mutants that bind TPP1 localize to telomeres but fail to repress a DNA damage response and inappropriate repair by A-NHEJ. Our results reveal that POT1 C terminus is essential to prevent initiation of genome instability permissive for tumorigenesis.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



Similar articles
-
Structural and functional analysis of the human POT1-TPP1 telomeric complex.Nat Commun. 2017 Apr 10;8:14928. doi: 10.1038/ncomms14928. Nat Commun. 2017. PMID: 28393830 Free PMC article.
-
TPP1 is a homologue of ciliate TEBP-beta and interacts with POT1 to recruit telomerase.Nature. 2007 Feb 1;445(7127):559-62. doi: 10.1038/nature05469. Epub 2007 Jan 21. Nature. 2007. PMID: 17237767
-
POT1-TPP1 Binding and Unfolding of Telomere DNA Discriminates against Structural Polymorphism.J Mol Biol. 2016 Jul 3;428(13):2695-708. doi: 10.1016/j.jmb.2016.04.031. Epub 2016 May 10. J Mol Biol. 2016. PMID: 27173378 Free PMC article.
-
[Advance in research on the function of telomeric shelterin component TPP1 and its relationship with characteristics of tumors].Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2016 Aug;33(4):573-7. doi: 10.3760/cma.j.issn.1003-9406.2016.04.033. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2016. PMID: 27455024 Review. Chinese.
-
Shelterin proteins and cancer.Asian Pac J Cancer Prev. 2015;16(8):3085-90. doi: 10.7314/apjcp.2015.16.8.3085. Asian Pac J Cancer Prev. 2015. PMID: 25921101 Review.
Cited by
-
TPP1 mutagenesis screens unravel shelterin interfaces and functions in hematopoiesis.JCI Insight. 2021 May 10;6(9):e138059. doi: 10.1172/jci.insight.138059. JCI Insight. 2021. PMID: 33822766 Free PMC article.
-
Initiation of Pulmonary Fibrosis after Silica Inhalation in Rats is linked with Dysfunctional Shelterin Complex and DNA Damage Response.Sci Rep. 2019 Jan 24;9(1):471. doi: 10.1038/s41598-018-36712-6. Sci Rep. 2019. PMID: 30679488 Free PMC article.
-
TINF2 is a major susceptibility gene in Danish patients with multiple primary melanoma.HGG Adv. 2023 Jul 23;4(4):100225. doi: 10.1016/j.xhgg.2023.100225. eCollection 2023 Oct 12. HGG Adv. 2023. PMID: 37646013 Free PMC article.
-
Evolving Linear Chromosomes and Telomeres: A C-Strand-Centric View.Trends Biochem Sci. 2018 May;43(5):314-326. doi: 10.1016/j.tibs.2018.02.008. Epub 2018 Mar 14. Trends Biochem Sci. 2018. PMID: 29550242 Free PMC article. Review.
-
Human shelterin protein POT1 prevents severe telomere instability induced by homology-directed DNA repair.EMBO J. 2020 Dec 1;39(23):e104500. doi: 10.15252/embj.2020104500. Epub 2020 Oct 19. EMBO J. 2020. PMID: 33073402 Free PMC article.
References
-
- Cech T. R. Beginning to understand the end of the chromosome. Cell 116, 273–279 (2004). - PubMed
-
- Makarov V. L., Hirose Y. & Langmore J. P. Long G tails at both ends of human chromosomes suggest a C strand degradation mechanism for telomere shortening. Cell 88, 657–666 (1997). - PubMed
-
- Baumann P. & Cech T. R. Pot1, the putative telomere end-binding protein in fission yeast and humans. Science 292, 1171–1175 (2001). - PubMed
-
- de Lange T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

