New perspective in diagnostics of mitochondrial disorders: two years' experience with whole-exome sequencing at a national paediatric centre
- PMID: 27290639
- PMCID: PMC4903158
- DOI: 10.1186/s12967-016-0930-9
New perspective in diagnostics of mitochondrial disorders: two years' experience with whole-exome sequencing at a national paediatric centre
Abstract
Background: Whole-exome sequencing (WES) has led to an exponential increase in identification of causative variants in mitochondrial disorders (MD).
Methods: We performed WES in 113 MD suspected patients from Polish paediatric reference centre, in whom routine testing failed to identify a molecular defect. WES was performed using TruSeqExome enrichment, followed by variant prioritization, validation by Sanger sequencing, and segregation with the disease phenotype in the family.
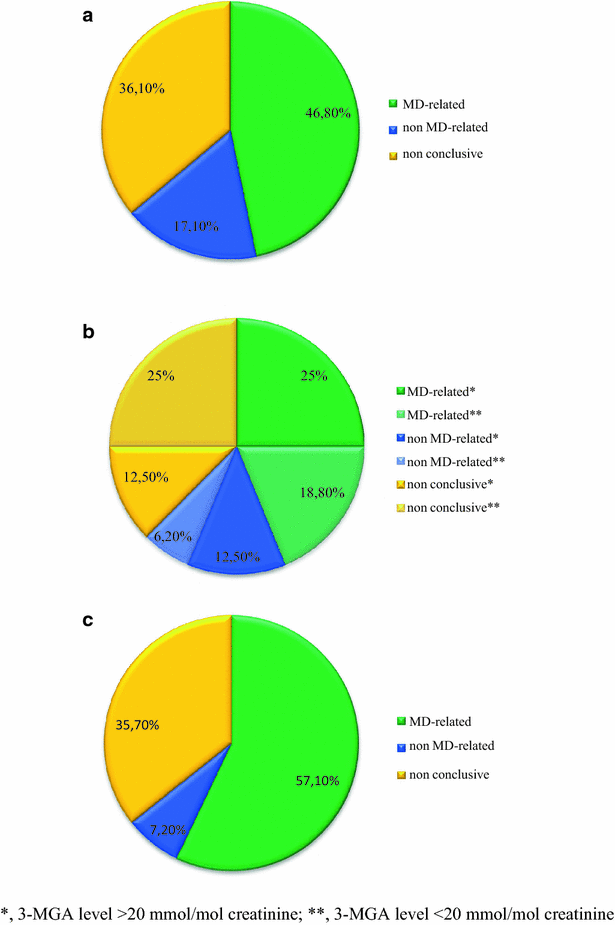
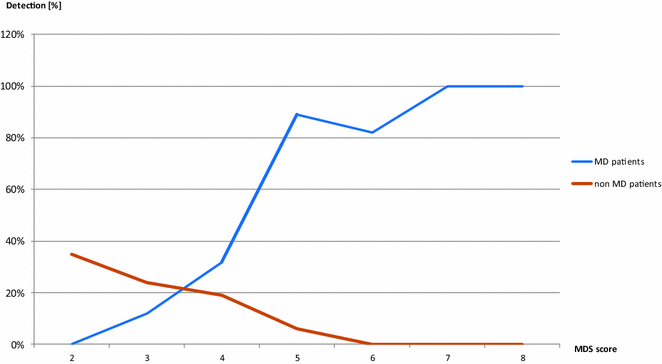
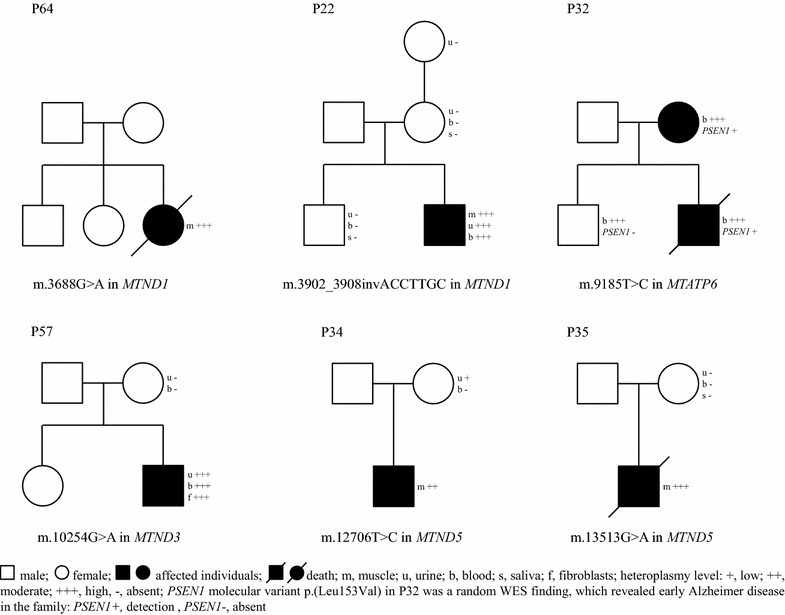
Results: Likely causative mutations were identified in 67 (59.3 %) patients; these included variants in mtDNA (6 patients) and nDNA: X-linked (9 patients), autosomal dominant (5 patients), and autosomal recessive (47 patients, 11 homozygotes). Novel variants accounted for 50.5 % (50/99) of all detected changes. In 47 patients, changes in 31 MD-related genes (ACAD9, ADCK3, AIFM1, CLPB, COX10, DLD, EARS2, FBXL4, MTATP6, MTFMT, MTND1, MTND3, MTND5, NAXE, NDUFS6, NDUFS7, NDUFV1, OPA1, PARS2, PC, PDHA1, POLG, RARS2, RRM2B, SCO2, SERAC1, SLC19A3, SLC25A12, TAZ, TMEM126B, VARS2) were identified. The ACAD9, CLPB, FBXL4, PDHA1 genes recurred more than twice suggesting higher general/ethnic prevalence. In 19 cases, variants in 18 non-MD related genes (ADAR, CACNA1A, CDKL5, CLN3, CPS1, DMD, DYSF, GBE1, GFAP, HSD17B4, MECP2, MYBPC3, PEX5, PGAP2, PIGN, PRF1, SBDS, SCN2A) were found. The percentage of positive WES results rose gradually with increasing probability of MD according to the Mitochondrial Disease Criteria (MDC) scale (from 36 to 90 % for low and high probability, respectively). The percentage of detected MD-related genes compared with non MD-related genes also grew with the increasing MD likelihood (from 20 to 97 %). Molecular diagnosis was established in 30/47 (63.8 %) neonates and in 17/28 (60.7 %) patients with basal ganglia involvement. Mutations in CLPB, SERAC1, TAZ genes were identified in neonates with 3-methylglutaconic aciduria (3-MGA) as a discriminative feature. New MD-related candidate gene (NDUFB8) is under verification.
Conclusions: We suggest WES rather than targeted NGS as the method of choice in diagnostics of MD in children, including neonates with 3-MGA aciduria, who died without determination of disease cause and with limited availability of laboratory data. There is a strong correlation between the degree of MD diagnosis by WES and MD likelihood expressed by the MDC scale.
Keywords: 3-methylglutaconic aciduria; Basal ganglia involvement; Candidate gene; Leigh syndrome; Mitochondrial disease criteria scale; Mitochondrial disorders; Neonates; Novel mutation; Whole-exome sequencing.
Figures
Similar articles
-
The utility of next-generation sequencing technologies in diagnosis of Mendelian mitochondrial diseases and reflections on clinical spectrum.J Pediatr Endocrinol Metab. 2021 Feb 24;34(4):417-430. doi: 10.1515/jpem-2020-0410. Print 2021 Apr 27. J Pediatr Endocrinol Metab. 2021. PMID: 33629572
-
Whole exome sequencing diagnosis of inborn errors of metabolism and other disorders in United Arab Emirates.Orphanet J Rare Dis. 2016 Jul 8;11(1):94. doi: 10.1186/s13023-016-0474-3. Orphanet J Rare Dis. 2016. PMID: 27391121 Free PMC article.
-
Bi-allelic CLPB mutations cause cataract, renal cysts, nephrocalcinosis and 3-methylglutaconic aciduria, a novel disorder of mitochondrial protein disaggregation.J Inherit Metab Dis. 2015 Mar;38(2):211-9. doi: 10.1007/s10545-015-9813-0. Epub 2015 Jan 18. J Inherit Metab Dis. 2015. PMID: 25595726
-
Translational Metabolism: A multidisciplinary approach towards precision diagnosis of inborn errors of metabolism in the omics era.J Inherit Metab Dis. 2019 Mar;42(2):197-208. doi: 10.1002/jimd.12008. Epub 2019 Feb 5. J Inherit Metab Dis. 2019. PMID: 30723938 Review.
-
Clinical manifestations and enzymatic activities of mitochondrial respiratory chain complexes in Pearson marrow-pancreas syndrome with 3-methylglutaconic aciduria: a case report and literature review.Eur J Pediatr. 2015 Dec;174(12):1593-602. doi: 10.1007/s00431-015-2576-7. Epub 2015 Jun 16. Eur J Pediatr. 2015. PMID: 26074369 Review.
Cited by
-
Facilitations and Hurdles of Genetic Testing in Neuromuscular Disorders.Diagnostics (Basel). 2021 Apr 14;11(4):701. doi: 10.3390/diagnostics11040701. Diagnostics (Basel). 2021. PMID: 33919863 Free PMC article. Review.
-
Analysis of compound heterozygous and homozygous mutations found in peripheral subunits of human respiratory Complex I, NDUFS1, NDUFS2, NDUFS8 and NDUFV1, by modeling in the E. coli enzyme.Mitochondrion. 2023 Jan;68:87-104. doi: 10.1016/j.mito.2022.11.007. Epub 2022 Nov 30. Mitochondrion. 2023. PMID: 36462614 Free PMC article.
-
NAXE deficiency: A neurometabolic disorder of NAD(P)HX repair amenable for metabolic correction.Mol Genet Metab. 2022 Jun;136(2):101-110. doi: 10.1016/j.ymgme.2022.04.003. Epub 2022 Apr 18. Mol Genet Metab. 2022. PMID: 35637064 Free PMC article. Review.
-
NDUFS6 related Leigh syndrome: a case report and review of the literature.J Hum Genet. 2019 Jul;64(7):637-645. doi: 10.1038/s10038-019-0594-4. Epub 2019 Apr 4. J Hum Genet. 2019. PMID: 30948790 Review.
-
FBXL4-Related Mitochondrial DNA Depletion Syndrome 13 (MTDPS13): A Case Report With a Comprehensive Mutation Review.Front Genet. 2019 Feb 5;10:39. doi: 10.3389/fgene.2019.00039. eCollection 2019. Front Genet. 2019. PMID: 30804983 Free PMC article.
References
-
- Shamseldin HE, Alshammari M, Al-Sheddi T, Salih MA, Alkhalidi H, Kentab A, Repetto GM, Hashem M, Alkuraya FS. Genomic analysis of mitochondrial diseases in a consanguineous population reveals novel candidate disease genes. J Med Genet. 2012;494:234–241. doi: 10.1136/jmedgenet-2012-100836. - DOI - PubMed
-
- Calvo SE, Compton AG, Hershman SG, Lim SC, Lieber DS, Tucker EJ, Laskowski A, Garone C, Liu S, Jaffe DB, Christodoulou J, Fletcher JM, Bruno DL, Goldblatt J, Dimauro S, Thorburn DR, Mootha VK. Molecular diagnosis of infantile mitochondrial disease with targeted next-generation sequencing. Sci Transl Med. 2012;4118:118ra10. - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

