SETD2: an epigenetic modifier with tumor suppressor functionality
- PMID: 27191891
- PMCID: PMC5226616
- DOI: 10.18632/oncotarget.9368
SETD2: an epigenetic modifier with tumor suppressor functionality
Abstract
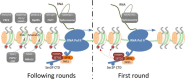
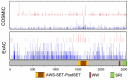
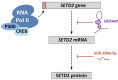
In the past decade important progress has been made in our understanding of the epigenetic regulatory machinery. It has become clear that genetic aberrations in multiple epigenetic modifier proteins are associated with various types of cancer. Moreover, targeting the epigenome has emerged as a novel tool to treat cancer patients. Recently, the first drugs have been reported that specifically target SETD2-negative tumors. In this review we discuss the studies on the associated protein, Set domain containing 2 (SETD2), a histone modifier for which mutations have only recently been associated with cancer development. Our review starts with the structural characteristics of SETD2 and extends to its corresponding function by combining studies on SETD2 function in yeast, Drosophila, Caenorhabditis elegans, mice, and humans. SETD2 is now generally known as the single human gene responsible for trimethylation of lysine 36 of Histone H3 (H3K36). H3K36me3 readers that recruit protein complexes to carry out specific processes, including transcription elongation, RNA processing, and DNA repair, determine the impact of this histone modification. Finally, we describe the prevalence of SETD2-inactivating mutations in cancer, with the highest frequency in clear cell Renal Cell Cancer, and explore how SETD2-inactivation might contribute to tumor development.
Keywords: H3K36me3; SETD2; ccRCC; histone modification; tumor suppressor gene.
Conflict of interest statement
There is no conflict of interest for any of the authors.
Figures
Similar articles
-
SETD2, an epigenetic tumor suppressor: a focused review on GI tumor.Front Biosci (Landmark Ed). 2020 Jan 1;25(4):781-797. doi: 10.2741/4834. Front Biosci (Landmark Ed). 2020. PMID: 31585917 Review.
-
The Benzene Hematotoxic and Reactive Metabolite 1,4-Benzoquinone Impairs the Activity of the Histone Methyltransferase SET Domain Containing 2 (SETD2) and Causes Aberrant Histone H3 Lysine 36 Trimethylation (H3K36me3).Mol Pharmacol. 2021 Sep;100(3):283-294. doi: 10.1124/molpharm.121.000303. Epub 2021 Jul 15. Mol Pharmacol. 2021. PMID: 34266924
-
Structure/Function Analysis of Recurrent Mutations in SETD2 Protein Reveals a Critical and Conserved Role for a SET Domain Residue in Maintaining Protein Stability and Histone H3 Lys-36 Trimethylation.J Biol Chem. 2016 Sep 30;291(40):21283-21295. doi: 10.1074/jbc.M116.739375. Epub 2016 Aug 15. J Biol Chem. 2016. PMID: 27528607 Free PMC article.
-
SETD2-dependent H3K36me3 plays a critical role in epigenetic regulation of the HPV31 life cycle.PLoS Pathog. 2018 Oct 12;14(10):e1007367. doi: 10.1371/journal.ppat.1007367. eCollection 2018 Oct. PLoS Pathog. 2018. PMID: 30312361 Free PMC article.
-
SETD2 as a regulator of N6-methyladenosine RNA methylation and modifiers in cancer.Eur J Cancer Prev. 2020 Nov;29(6):556-564. doi: 10.1097/CEJ.0000000000000587. Eur J Cancer Prev. 2020. PMID: 33021769 Review.
Cited by
-
Autophagy regulation by RNA alternative splicing and implications in human diseases.Nat Commun. 2022 May 18;13(1):2735. doi: 10.1038/s41467-022-30433-1. Nat Commun. 2022. PMID: 35585060 Free PMC article. Review.
-
SETD2: from chromatin modifier to multipronged regulator of the genome and beyond.Cell Mol Life Sci. 2022 Jun 6;79(6):346. doi: 10.1007/s00018-022-04352-9. Cell Mol Life Sci. 2022. PMID: 35661267 Free PMC article. Review.
-
Genomic landscape of pancreatic neuroendocrine tumours: the International Cancer Genome Consortium.J Endocrinol. 2018 Mar;236(3):R161-R167. doi: 10.1530/JOE-17-0560. Epub 2018 Jan 10. J Endocrinol. 2018. PMID: 29321190 Free PMC article. Review.
-
Shaping the cellular landscape with Set2/SETD2 methylation.Cell Mol Life Sci. 2017 Sep;74(18):3317-3334. doi: 10.1007/s00018-017-2517-x. Epub 2017 Apr 6. Cell Mol Life Sci. 2017. PMID: 28386724 Free PMC article. Review.
-
Molecular Basis of Cisplatin Resistance in Testicular Germ Cell Tumors.Cancers (Basel). 2019 Sep 6;11(9):1316. doi: 10.3390/cancers11091316. Cancers (Basel). 2019. PMID: 31500094 Free PMC article.
References
-
- Faber PW, Barnes GT, Srinidhi J, Chen J, Gusella JF, MacDonald ME. Huntingtin interacts with a family of WW domain proteins. Hum Mol Genet. 1998;7:1463–74. - PubMed
-
- Mao M, Fu G, Wu JS, Zhang QH, Zhou J, Kan LX, Huang QH, He KL, Gu BW, Han ZG, Shen Y, Gu J, Yu YP, et al. Identification of genes expressed in human CD34 hematopoietic stem/progenitor cells by expressed sequence tags and efficient full-length cDNA cloning. Proc Natl Acad Sci USA. 1998;95:8175–80. - PMC - PubMed
-
- Zhang QH, Ye M, Wu XY, Ren SX, Zhao M, Zhao CJ, Fu G, Shen Y, Fan HY, Lu G, Zhong M, Xu XR, Han ZG, et al. Cloning and functional analysis of cDNAs with open reading frames for 300 previously undefined genes expressed in CD34 hematopoietic stem/progenitor cells. Genome Res. 2000;10:1546–60. - PMC - PubMed
-
- Sun X, Wei J, Wu X, Hu M, Wang L, Wang H, Zhang Q, Chen S, Huang Q, Chen Z. Identification and characterization of a novel human histone H3 lysine 36-specific methyltransferase. J Biol Chem. 2005;280:35261–71. - PubMed
-
- Rega S, Stiewe T, Chang D, Pollmeier B, Esche H, Bardenheuer W, Marquitan G, Pützer BM. Identification of the full-length huntingtin-interacting protein p231HBP/HYPB as a DNA-binding factor. Mol Cell Neurosci. 2001;18:68–79. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

