hnRNP K Is a Haploinsufficient Tumor Suppressor that Regulates Proliferation and Differentiation Programs in Hematologic Malignancies
- PMID: 26412324
- PMCID: PMC4652598
- DOI: 10.1016/j.ccell.2015.09.001
hnRNP K Is a Haploinsufficient Tumor Suppressor that Regulates Proliferation and Differentiation Programs in Hematologic Malignancies
Abstract
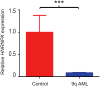
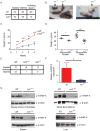
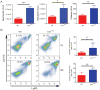
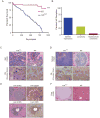
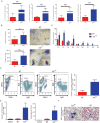
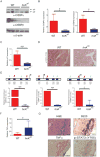
hnRNP K regulates cellular programs, and changes in its expression and mutational status have been implicated in neoplastic malignancies. To directly examine its role in tumorigenesis, we generated a mouse model harboring an Hnrnpk knockout allele (Hnrnpk(+/-)). Hnrnpk haploinsufficiency resulted in reduced survival, increased tumor formation, genomic instability, and the development of transplantable hematopoietic neoplasms with myeloproliferation. Reduced hnRNP K expression attenuated p21 activation, downregulated C/EBP levels, and activated STAT3 signaling. Additionally, analysis of samples from primary acute myeloid leukemia patients harboring a partial deletion of chromosome 9 revealed a significant decrease in HNRNPK expression. Together, these data implicate hnRNP K in the development of hematological disorders and suggest hnRNP K acts as a tumor suppressor.
Copyright © 2015 Elsevier Inc. All rights reserved.
Conflict of interest statement
Figures

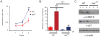
Similar articles
-
Aberrant hnRNP K expression: All roads lead to cancer.Cell Cycle. 2016 Jun 17;15(12):1552-7. doi: 10.1080/15384101.2016.1164372. Cell Cycle. 2016. PMID: 27049467 Free PMC article. Review.
-
Insight into the mechanism of AML del(9q) progression: hnRNP K targets the myeloid master regulators CEBPA (C/EBPα) and SPI1 (PU.1).Biochim Biophys Acta Gene Regul Mech. 2024 Mar;1867(1):195004. doi: 10.1016/j.bbagrm.2023.195004. Epub 2023 Nov 25. Biochim Biophys Acta Gene Regul Mech. 2024. PMID: 38008244
-
Clinicopathologic and molecular characterization of myeloid neoplasms harboring isochromosome 17(q10).Am J Hematol. 2014 Aug;89(8):862. doi: 10.1002/ajh.23755. Epub 2014 May 16. Am J Hematol. 2014. PMID: 24796269 No abstract available.
-
Limited miR-17-92 overexpression drives hematologic malignancies.Leuk Res. 2015 Mar;39(3):335-41. doi: 10.1016/j.leukres.2014.12.002. Epub 2014 Dec 10. Leuk Res. 2015. PMID: 25597017 Free PMC article.
-
Role of microRNAs in hematological malignancies.Expert Rev Hematol. 2009 Aug;2(4):415-23. doi: 10.1586/ehm.09.32. Expert Rev Hematol. 2009. PMID: 21082946 Review.
Cited by
-
Identification of heterogeneous nuclear ribonucleoprotein as a candidate biomarker for diagnosis and prognosis of hepatocellular carcinoma.J Gastrointest Oncol. 2021 Oct;12(5):2361-2376. doi: 10.21037/jgo-21-468. J Gastrointest Oncol. 2021. PMID: 34790398 Free PMC article.
-
Cancer the'RBP'eutics-RNA-binding proteins as therapeutic targets for cancer.Pharmacol Ther. 2019 Nov;203:107390. doi: 10.1016/j.pharmthera.2019.07.001. Epub 2019 Jul 11. Pharmacol Ther. 2019. PMID: 31302171 Free PMC article. Review.
-
Splicing-factor alterations in cancers.RNA. 2016 Sep;22(9):1285-301. doi: 10.1261/rna.057919.116. RNA. 2016. PMID: 27530828 Free PMC article. Review.
-
RNA splicing factors as oncoproteins and tumour suppressors.Nat Rev Cancer. 2016 Jul;16(7):413-30. doi: 10.1038/nrc.2016.51. Epub 2016 Jun 10. Nat Rev Cancer. 2016. PMID: 27282250 Free PMC article. Review.
-
Integrated nuclear proteomics and transcriptomics identifies S100A4 as a therapeutic target in acute myeloid leukemia.Leukemia. 2020 Feb;34(2):427-440. doi: 10.1038/s41375-019-0596-4. Epub 2019 Oct 14. Leukemia. 2020. PMID: 31611628 Free PMC article.
References
-
- Burnett A, Wetzler M, Lowenberg B. Therapeutic advances in acute myeloid leukemia. J Clin Oncol. 2011;29:487–494. - PubMed
-
- Enge M, Bao W, Hedstrom E, Jackson SP, Moumen A, Selivanova G. MDM2-dependent downregulation of p21 and hnRNP K provides a switch between apoptosis and growth arrest induced by pharmacologically activated p53. Cancer Cell. 2009;15:171–183. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

