ZBTB20 is a sequence-specific transcriptional repressor of alpha-fetoprotein gene
- PMID: 26173901
- PMCID: PMC4648434
- DOI: 10.1038/srep11979
ZBTB20 is a sequence-specific transcriptional repressor of alpha-fetoprotein gene
Abstract
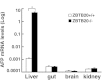
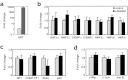
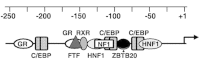
Alpha-fetoprotein (AFP) represents a classical model system to study developmental gene regulation in mammalian cells. We previously reported that liver ZBTB20 is developmentally regulated and plays a central role in AFP postnatal repression. Here we show that ZBTB20 is a sequence-specific transcriptional repressor of AFP. By ELISA-based DNA-protein binding assay and conventional gel shift assay, we successfully identified a ZBTB20-binding site at -104/-86 of mouse AFP gene, flanked by two HNF1 sites and two C/EBP sites in the proximal promoter. Importantly, mutation of the core sequence in this site fully abolished its binding to ZBTB20 in vitro, as well as the repression of AFP promoter activity by ZBTB20. The unique ZBTB20 site was highly conserved in rat and human AFP genes, but absent in albumin genes. These help to explain the autonomous regulation of albumin and AFP genes in the liver after birth. Furthermore, we demonstrated that transcriptional repression of AFP gene by ZBTB20 was liver-specific. ZBTB20 was dispensable for AFP silencing in other tissues outside liver. Our data define a cognate ZBTB20 site in AFP promoter which mediates the postnatal repression of AFP gene in the liver.
Figures


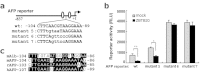


Similar articles
-
Zinc finger protein ZBTB20 is a key repressor of alpha-fetoprotein gene transcription in liver.Proc Natl Acad Sci U S A. 2008 Aug 5;105(31):10859-64. doi: 10.1073/pnas.0800647105. Epub 2008 Jul 31. Proc Natl Acad Sci U S A. 2008. PMID: 18669658 Free PMC article.
-
Functional analysis of developmentally regulated chromatin-hypersensitive domains carrying the alpha 1-fetoprotein gene promoter and the albumin/alpha 1-fetoprotein intergenic enhancer.Mol Cell Biol. 1993 Mar;13(3):1619-33. doi: 10.1128/mcb.13.3.1619-1633.1993. Mol Cell Biol. 1993. PMID: 7680097 Free PMC article.
-
Transcription factor interactions and chromatin modifications associated with p53-mediated, developmental repression of the alpha-fetoprotein gene.Mol Cell Biol. 2005 Mar;25(6):2147-57. doi: 10.1128/MCB.25.6.2147-2157.2005. Mol Cell Biol. 2005. PMID: 15743813 Free PMC article.
-
Zhx2 and Zbtb20: novel regulators of postnatal alpha-fetoprotein repression and their potential role in gene reactivation during liver cancer.Semin Cancer Biol. 2011 Feb;21(1):21-7. doi: 10.1016/j.semcancer.2011.01.001. Epub 2011 Jan 7. Semin Cancer Biol. 2011. PMID: 21216289 Free PMC article. Review.
-
Molecular mechanisms of alpha-fetoprotein gene expression.Biochemistry (Mosc). 2000 Jan;65(1):117-33. Biochemistry (Mosc). 2000. PMID: 10702646 Review.
Cited by
-
ZBTB20 is required for anterior pituitary development and lactotrope specification.Nat Commun. 2016 Apr 15;7:11121. doi: 10.1038/ncomms11121. Nat Commun. 2016. PMID: 27079169 Free PMC article.
-
Intracellular alpha-fetoprotein mitigates hepatocyte apoptosis and necroptosis by inhibiting endoplasmic reticulum stress.World J Gastroenterol. 2022 Jul 14;28(26):3201-3217. doi: 10.3748/wjg.v28.i26.3201. World J Gastroenterol. 2022. PMID: 36051342 Free PMC article.
-
Increased Expression of SETD7 Promotes Cell Proliferation by Regulating Cell Cycle and Indicates Poor Prognosis in Hepatocellular Carcinoma.PLoS One. 2016 May 16;11(5):e0154939. doi: 10.1371/journal.pone.0154939. eCollection 2016. PLoS One. 2016. PMID: 27183310 Free PMC article.
-
TUG1 Is a Regulator of AFP and Serves as Prognostic Marker in Non-Hepatitis B Non-Hepatitis C Hepatocellular Carcinoma.Cells. 2020 Jan 21;9(2):262. doi: 10.3390/cells9020262. Cells. 2020. PMID: 31973032 Free PMC article.
-
Regulation of hepatic lipogenesis by the zinc finger protein Zbtb20.Nat Commun. 2017 Mar 22;8:14824. doi: 10.1038/ncomms14824. Nat Commun. 2017. PMID: 28327662 Free PMC article.
References
-
- Tilghman S. M. The structure and regulation of the alpha-fetoprotein and albumin genes. Oxf Surv Eukaryot Genes 2, 160–206 (1985). - PubMed
-
- Spear B. T. Alpha-fetoprotein gene regulation: lessons from transgenic mice. Semin Cancer Biol 9, 109–116 (1999). - PubMed
-
- Vacher J. & Tilghman S. M. Dominant negative regulation of the mouse alpha-fetoprotein gene in adult liver. Science 250, 1732–1735 (1990). - PubMed
-
- Hammer R. E., Krumlauf R., Camper S.A., Brinster R.L. & Tilghman S.M. Diversity of alpha-fetoprotein gene expression in mice is generated by a combination of separate enhancer elements. Science 235, 53–58 (1987). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

