Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network
- PMID: 23911319
- PMCID: PMC3894107
- DOI: 10.1016/j.cell.2013.06.049
Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network
Abstract
Genes disrupted in schizophrenia may be revealed by de novo mutations in affected persons from otherwise healthy families. Furthermore, during normal brain development, genes are expressed in patterns specific to developmental stage and neuroanatomical structure. We identified de novo mutations in persons with schizophrenia and then mapped the responsible genes onto transcriptome profiles of normal human brain tissues from age 13 weeks gestation to adulthood. In the dorsolateral and ventrolateral prefrontal cortex during fetal development, genes harboring damaging de novo mutations in schizophrenia formed a network significantly enriched for transcriptional coexpression and protein interaction. The 50 genes in the network function in neuronal migration, synaptic transmission, signaling, transcriptional regulation, and transport. These results suggest that disruptions of fetal prefrontal cortical neurogenesis are critical to the pathophysiology of schizophrenia. These results also support the feasibility of integrating genomic and transcriptome analyses to map critical neurodevelopmental processes in time and space in the brain.
Copyright © 2013 Elsevier Inc. All rights reserved.
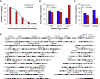
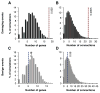
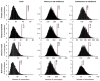
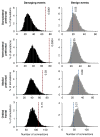
Figures


Similar articles
-
De novo mutations in schizophrenia disrupt genes co-expressed in fetal prefrontal cortex.Neuropsychopharmacology. 2014 Jan;39(1):238-9. doi: 10.1038/npp.2013.219. Neuropsychopharmacology. 2014. PMID: 24317315 Free PMC article. No abstract available.
-
Gene expression profile associated with postnatal development of pyramidal neurons in the human prefrontal cortex implicates ubiquitin ligase E3 in the pathophysiology of schizophrenia onset.J Psychiatr Res. 2018 Jul;102:110-117. doi: 10.1016/j.jpsychires.2018.03.013. Epub 2018 Apr 3. J Psychiatr Res. 2018. PMID: 29635114 Free PMC article.
-
Prefrontal Coexpression of Schizophrenia Risk Genes Is Associated With Treatment Response in Patients.Biol Psychiatry. 2019 Jul 1;86(1):45-55. doi: 10.1016/j.biopsych.2019.03.981. Epub 2019 Apr 3. Biol Psychiatry. 2019. PMID: 31126695
-
Specific relationship between prefrontal neuronal N-acetylaspartate and activation of the working memory cortical network in schizophrenia.Am J Psychiatry. 2000 Jan;157(1):26-33. doi: 10.1176/ajp.157.1.26. Am J Psychiatry. 2000. PMID: 10618009 Review.
-
Schizophrenia: a tale of two critical periods for prefrontal cortical development.Transl Psychiatry. 2015 Aug 18;5(8):e623. doi: 10.1038/tp.2015.115. Transl Psychiatry. 2015. PMID: 26285133 Free PMC article. Review.
Cited by
-
Genetics of schizophrenia (Review).Exp Ther Med. 2020 Oct;20(4):3462-3468. doi: 10.3892/etm.2020.8973. Epub 2020 Jul 7. Exp Ther Med. 2020. PMID: 32905096 Free PMC article. Review.
-
2020 William Allan Award introduction: Mary-Claire King.Am J Hum Genet. 2021 Mar 4;108(3):383-385. doi: 10.1016/j.ajhg.2020.12.011. Am J Hum Genet. 2021. PMID: 33667390 Free PMC article.
-
The subventricular zone in the immature piglet brain: anatomy and exodus of neuroblasts into white matter after traumatic brain injury.Dev Neurosci. 2015;37(2):115-30. doi: 10.1159/000369091. Epub 2015 Feb 10. Dev Neurosci. 2015. PMID: 25678047 Free PMC article.
-
Investigation of Schizophrenia with Human Induced Pluripotent Stem Cells.Adv Neurobiol. 2020;25:155-206. doi: 10.1007/978-3-030-45493-7_6. Adv Neurobiol. 2020. PMID: 32578147 Free PMC article. Review.
-
Germline ETV6 mutations in familial thrombocytopenia and hematologic malignancy.Nat Genet. 2015 Feb;47(2):180-5. doi: 10.1038/ng.3177. Epub 2015 Jan 12. Nat Genet. 2015. PMID: 25581430 Free PMC article.
References
-
- Anton ES, Kreidberg JA, Rakic P. Distinct functions of alpha3 and alpha(v) integrin receptors in neuronal migration and laminar organization of the cerebral cortex. Neuron. 1999;22:277–289. - PubMed
-
- Barabási AL, Oltvai ZN. Network biology: understanding the cell’s functional organization. Nat Rev Genet. 2004;5:101–113. - PubMed
-
- BrainSpan: Atlas of the Developing Human Brain. 2013 www.brainspan.org.
Publication types
MeSH terms
Grants and funding
- R03TW008696/TW/FIC NIH HHS/United States
- R01MH083989/MH/NIMH NIH HHS/United States
- R01 MH083849/MH/NIMH NIH HHS/United States
- R01 MH083989/MH/NIMH NIH HHS/United States
- R01MH083849/MH/NIMH NIH HHS/United States
- R01MH084071/MH/NIMH NIH HHS/United States
- R03 TW008696/TW/FIC NIH HHS/United States
- R01MH083756/MH/NIMH NIH HHS/United States
- R01 MH083756/MH/NIMH NIH HHS/United States
- R01 MH086135/MH/NIMH NIH HHS/United States
- R01 MH084071/MH/NIMH NIH HHS/United States
- U01 MH096844/MH/NIMH NIH HHS/United States
- R01 MH093533/MH/NIMH NIH HHS/United States
- R01 CA157744/CA/NCI NIH HHS/United States
- T32 GM007266/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

