Intrinsic protein-protein interaction-mediated and chaperonin-assisted sequential assembly of stable bardet-biedl syndrome protein complex, the BBSome
- PMID: 22500027
- PMCID: PMC3370246
- DOI: 10.1074/jbc.M112.341487
Intrinsic protein-protein interaction-mediated and chaperonin-assisted sequential assembly of stable bardet-biedl syndrome protein complex, the BBSome
Abstract
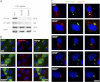
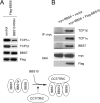
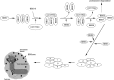
The pleiotropic features of obesity, retinal degeneration, polydactyly, kidney abnormalities, cognitive impairment, hypertension, and diabetes found in Bardet-Biedl syndrome (BBS) make this disorder an important model disorder for identifying molecular mechanisms involved in common human diseases. To date, 16 BBS genes have been reported, seven of which (BBS1, 2, 4, 5, 7, 8, and 9) code for proteins that form a complex known as the BBSome. The function of the BBSome involves ciliary membrane biogenesis. Three additional BBS genes (BBS6, BBS10, and BBS12) have homology to type II chaperonins and interact with CCT/TRiC proteins and BBS7 to form a complex termed the BBS-chaperonin complex. This complex is required for BBSome assembly. Little is known about the process and the regulation of BBSome formation. We utilized point mutations and null alleles of BBS proteins to disrupt assembly of the BBSome leading to the accumulation of BBSome assembly intermediates. By characterizing BBSome assembly intermediates, we show that the BBS-chaperonin complex plays a role in BBS7 stability. BBS7 interacts with BBS2 and becomes part of a BBS7-BBS2-BBS9 assembly intermediate referred to as the BBSome core complex because it forms the core of the BBSome. BBS1, BBS5, BBS8, and finally BBS4 are added to the BBSome core to form the complete BBSome.
Figures





Similar articles
-
BBS6, BBS10, and BBS12 form a complex with CCT/TRiC family chaperonins and mediate BBSome assembly.Proc Natl Acad Sci U S A. 2010 Jan 26;107(4):1488-93. doi: 10.1073/pnas.0910268107. Epub 2010 Jan 4. Proc Natl Acad Sci U S A. 2010. PMID: 20080638 Free PMC article.
-
Mutations in chaperonin-like BBS genes are a major contributor to disease development in a multiethnic Bardet-Biedl syndrome patient population.J Med Genet. 2010 Jul;47(7):453-63. doi: 10.1136/jmg.2009.073205. Epub 2010 May 14. J Med Genet. 2010. PMID: 20472660
-
BBS7 is required for BBSome formation and its absence in mice results in Bardet-Biedl syndrome phenotypes and selective abnormalities in membrane protein trafficking.J Cell Sci. 2013 Jun 1;126(Pt 11):2372-80. doi: 10.1242/jcs.111740. Epub 2013 Apr 9. J Cell Sci. 2013. PMID: 23572516 Free PMC article.
-
Bardet-Biedl syndrome: The pleiotropic role of the chaperonin-like BBS6, 10, and 12 proteins.Am J Med Genet C Semin Med Genet. 2022 Mar;190(1):9-19. doi: 10.1002/ajmg.c.31970. Epub 2022 Apr 4. Am J Med Genet C Semin Med Genet. 2022. PMID: 35373910 Free PMC article. Review.
-
Bardet-Biedl Syndrome as a Chaperonopathy: Dissecting the Major Role of Chaperonin-Like BBS Proteins (BBS6-BBS10-BBS12).Front Mol Biosci. 2017 Jul 31;4:55. doi: 10.3389/fmolb.2017.00055. eCollection 2017. Front Mol Biosci. 2017. PMID: 28824921 Free PMC article. Review.
Cited by
-
Correction of cilia structure and function alleviates multi-organ pathology in Bardet-Biedl syndrome mice.Hum Mol Genet. 2020 Aug 29;29(15):2508-2522. doi: 10.1093/hmg/ddaa138. Hum Mol Genet. 2020. PMID: 32620959 Free PMC article.
-
Primary Cilium, An Unsung Hero in Maintaining Functional β-cell Population.Yale J Biol Med. 2019 Sep 20;92(3):471-480. eCollection 2019 Sep. Yale J Biol Med. 2019. PMID: 31543709 Free PMC article. Review.
-
Exome sequencing of Bardet-Biedl syndrome patient identifies a null mutation in the BBSome subunit BBIP1 (BBS18).J Med Genet. 2014 Feb;51(2):132-6. doi: 10.1136/jmedgenet-2013-101785. Epub 2013 Sep 11. J Med Genet. 2014. PMID: 24026985 Free PMC article.
-
Centriolar satellites: key mediators of centrosome functions.Cell Mol Life Sci. 2015 Jan;72(1):11-23. doi: 10.1007/s00018-014-1711-3. Epub 2014 Aug 31. Cell Mol Life Sci. 2015. PMID: 25173771 Free PMC article. Review.
-
Direct evidence for BBSome-associated intraflagellar transport reveals distinct properties of native mammalian cilia.Nat Commun. 2014 Dec 15;5:5813. doi: 10.1038/ncomms6813. Nat Commun. 2014. PMID: 25504142 Free PMC article.
References
-
- Lucker B. F., Behal R. H., Qin H., Siron L. C., Taggart W. D., Rosenbaum J. L., Cole D. G. (2005) Characterization of the intraflagellar transport complex B core. Direct interaction of the IFT81 and IFT74/72 subunits. J. Biol. Chem. 280, 27688–27696 - PubMed
-
- Oeffinger M., Wei K. E., Rogers R., DeGrasse J. A., Chait B. T., Aitchison J. D., Rout M. P. (2007) Comprehensive analysis of diverse ribonucleoprotein complexes. Nat. Methods 4, 951–956 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

