Patterns and rates of exonic de novo mutations in autism spectrum disorders
- PMID: 22495311
- PMCID: PMC3613847
- DOI: 10.1038/nature11011
Patterns and rates of exonic de novo mutations in autism spectrum disorders
Abstract
Autism spectrum disorders (ASD) are believed to have genetic and environmental origins, yet in only a modest fraction of individuals can specific causes be identified. To identify further genetic risk factors, here we assess the role of de novo mutations in ASD by sequencing the exomes of ASD cases and their parents (n = 175 trios). Fewer than half of the cases (46.3%) carry a missense or nonsense de novo variant, and the overall rate of mutation is only modestly higher than the expected rate. In contrast, the proteins encoded by genes that harboured de novo missense or nonsense mutations showed a higher degree of connectivity among themselves and to previous ASD genes as indexed by protein-protein interaction screens. The small increase in the rate of de novo events, when taken together with the protein interaction results, are consistent with an important but limited role for de novo point mutations in ASD, similar to that documented for de novo copy number variants. Genetic models incorporating these data indicate that most of the observed de novo events are unconnected to ASD; those that do confer risk are distributed across many genes and are incompletely penetrant (that is, not necessarily sufficient for disease). Our results support polygenic models in which spontaneous coding mutations in any of a large number of genes increases risk by 5- to 20-fold. Despite the challenge posed by such models, results from de novo events and a large parallel case-control study provide strong evidence in favour of CHD8 and KATNAL2 as genuine autism risk factors.
Figures
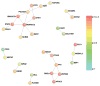

Comment in
-
Neurogenetics: Unravelling the genetics of autism.Nat Rev Neurosci. 2012 May 10;13(6):359. doi: 10.1038/nrn3259. Nat Rev Neurosci. 2012. PMID: 22573024 No abstract available.
-
Human genetics: Fruits of exome sequencing for autism.Nat Rev Genet. 2012 May 15;13(6):377. doi: 10.1038/nrg3248. Nat Rev Genet. 2012. PMID: 22585064 No abstract available.
Similar articles
-
Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations.Nature. 2012 Apr 4;485(7397):246-50. doi: 10.1038/nature10989. Nature. 2012. PMID: 22495309 Free PMC article.
-
De novo mutations revealed by whole-exome sequencing are strongly associated with autism.Nature. 2012 Apr 4;485(7397):237-41. doi: 10.1038/nature10945. Nature. 2012. PMID: 22495306 Free PMC article.
-
Meta-Analyses Support Previous and Novel Autism Candidate Genes: Outcomes of an Unexplored Brazilian Cohort.Autism Res. 2020 Feb;13(2):199-206. doi: 10.1002/aur.2238. Epub 2019 Nov 6. Autism Res. 2020. PMID: 31696658
-
The genetics of autism.Pediatrics. 2004 May;113(5):e472-86. doi: 10.1542/peds.113.5.e472. Pediatrics. 2004. PMID: 15121991 Review.
-
A de novo convergence of autism genetics and molecular neuroscience.Trends Neurosci. 2014 Feb;37(2):95-105. doi: 10.1016/j.tins.2013.11.005. Epub 2013 Dec 30. Trends Neurosci. 2014. PMID: 24387789 Free PMC article. Review.
Cited by
-
Genetic Influences on Cognitive Dysfunction in Schizophrenia.Curr Top Behav Neurosci. 2023;63:291-314. doi: 10.1007/7854_2022_388. Curr Top Behav Neurosci. 2023. PMID: 36029459
-
Sex differences in autism spectrum disorders.Curr Opin Neurol. 2013 Apr;26(2):146-53. doi: 10.1097/WCO.0b013e32835ee548. Curr Opin Neurol. 2013. PMID: 23406909 Free PMC article. Review.
-
Discovery of Rare Mutations in Autism: Elucidating Neurodevelopmental Mechanisms.Neurotherapeutics. 2015 Jul;12(3):553-71. doi: 10.1007/s13311-015-0363-9. Neurotherapeutics. 2015. PMID: 26105128 Free PMC article. Review.
-
Analysis of the chromosome X exome in patients with autism spectrum disorders identified novel candidate genes, including TMLHE.Transl Psychiatry. 2012 Oct 23;2(10):e179. doi: 10.1038/tp.2012.102. Transl Psychiatry. 2012. PMID: 23092983 Free PMC article.
-
HYST: a hybrid set-based test for genome-wide association studies, with application to protein-protein interaction-based association analysis.Am J Hum Genet. 2012 Sep 7;91(3):478-88. doi: 10.1016/j.ajhg.2012.08.004. Am J Hum Genet. 2012. PMID: 22958900 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- R01 MH089208/MH/NIMH NIH HHS/United States
- U54 HG003067/HG/NHGRI NIH HHS/United States
- P50 GM071558/GM/NIGMS NIH HHS/United States
- R01 MH089004/MH/NIMH NIH HHS/United States
- R01 MH061009/MH/NIMH NIH HHS/United States
- R01 MH057881/MH/NIMH NIH HHS/United States
- UL1 RR024975/RR/NCRR NIH HHS/United States
- R01 MH089482/MH/NIMH NIH HHS/United States
- R01MH084676/MH/NIMH NIH HHS/United States
- TL1 RR024978/RR/NCRR NIH HHS/United States
- P30 HD015052/HD/NICHD NIH HHS/United States
- KL2 RR024977/RR/NCRR NIH HHS/United States
- T32 GM007753/GM/NIGMS NIH HHS/United States
- U54 HG003273/HG/NHGRI NIH HHS/United States
- R01MH089175/MH/NIMH NIH HHS/United States
- P50 HD055751/HD/NICHD NIH HHS/United States
- R01 MH089025/MH/NIMH NIH HHS/United States
- R01MH089208/MH/NIMH NIH HHS/United States
- R01 MH089175/MH/NIMH NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

