Mutations in axonemal dynein assembly factor DNAAF3 cause primary ciliary dyskinesia
- PMID: 22387996
- PMCID: PMC3315610
- DOI: 10.1038/ng.1106
Mutations in axonemal dynein assembly factor DNAAF3 cause primary ciliary dyskinesia
Abstract
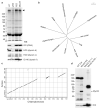
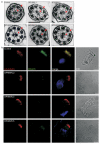
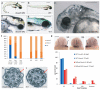
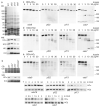
Primary ciliary dyskinesia most often arises from loss of the dynein motors that power ciliary beating. Here we show that DNAAF3 (also known as PF22), a previously uncharacterized protein, is essential for the preassembly of dyneins into complexes before their transport into cilia. We identified loss-of-function mutations in the human DNAAF3 gene in individuals from families with situs inversus and defects in the assembly of inner and outer dynein arms. Knockdown of dnaaf3 in zebrafish likewise disrupts dynein arm assembly and ciliary motility, causing primary ciliary dyskinesia phenotypes that include hydrocephalus and laterality malformations. Chlamydomonas reinhardtii PF22 is exclusively cytoplasmic, and a PF22-null mutant cannot assemble any outer and some inner dynein arms. Altered abundance of dynein subunits in mutant cytoplasm suggests that DNAAF3 (PF22) acts at a similar stage as other preassembly proteins, for example, DNAAF2 (also known as PF13 or KTU) and DNAAF1 (also known as ODA7 or LRRC50), in the dynein preassembly pathway. These results support the existence of a conserved, multistep pathway for the cytoplasmic formation of assembly competent ciliary dynein complexes.
Figures


Similar articles
-
Loss-of-function mutations in the human ortholog of Chlamydomonas reinhardtii ODA7 disrupt dynein arm assembly and cause primary ciliary dyskinesia.Am J Hum Genet. 2009 Dec;85(6):890-6. doi: 10.1016/j.ajhg.2009.11.008. Am J Hum Genet. 2009. PMID: 19944405 Free PMC article.
-
CCDC151 mutations cause primary ciliary dyskinesia by disruption of the outer dynein arm docking complex formation.Am J Hum Genet. 2014 Sep 4;95(3):257-74. doi: 10.1016/j.ajhg.2014.08.005. Am J Hum Genet. 2014. PMID: 25192045 Free PMC article.
-
Methylation of ciliary dynein motors involves the essential cytosolic assembly factor DNAAF3/PF22.Proc Natl Acad Sci U S A. 2024 Jan 30;121(5):e2318522121. doi: 10.1073/pnas.2318522121. Epub 2024 Jan 23. Proc Natl Acad Sci U S A. 2024. PMID: 38261620 Free PMC article.
-
Primary Ciliary Dyskinesia.2007 Jan 24 [updated 2019 Dec 5]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. 2007 Jan 24 [updated 2019 Dec 5]. In: Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Amemiya A, editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle; 1993–2025. PMID: 20301301 Free Books & Documents. Review.
-
Ciliary motility: the components and cytoplasmic preassembly mechanisms of the axonemal dyneins.Differentiation. 2012 Feb;83(2):S23-9. doi: 10.1016/j.diff.2011.11.009. Epub 2011 Dec 7. Differentiation. 2012. PMID: 22154137 Review.
Cited by
-
The ciliopathy-associated CPLANE proteins direct basal body recruitment of intraflagellar transport machinery.Nat Genet. 2016 Jun;48(6):648-56. doi: 10.1038/ng.3558. Epub 2016 May 9. Nat Genet. 2016. PMID: 27158779 Free PMC article.
-
Congenital hydrocephalus: a review of recent advances in genetic etiology and molecular mechanisms.Mil Med Res. 2024 Aug 12;11(1):54. doi: 10.1186/s40779-024-00560-5. Mil Med Res. 2024. PMID: 39135208 Free PMC article. Review.
-
Primary ciliary dyskinesia, an orphan disease.Eur J Pediatr. 2013 Feb;172(2):151-62. doi: 10.1007/s00431-012-1785-6. Epub 2012 Jul 10. Eur J Pediatr. 2013. PMID: 22777640 Review.
-
Splice-site mutations in the axonemal outer dynein arm docking complex gene CCDC114 cause primary ciliary dyskinesia.Am J Hum Genet. 2013 Jan 10;92(1):88-98. doi: 10.1016/j.ajhg.2012.11.002. Epub 2012 Dec 20. Am J Hum Genet. 2013. PMID: 23261303 Free PMC article.
-
Diagnosis and management of primary ciliary dyskinesia.Cilia. 2015 Jan 22;4(1):2. doi: 10.1186/s13630-014-0011-8. eCollection 2015. Cilia. 2015. PMID: 25610612 Free PMC article.
References
-
- Coren ME, Meeks M, Morrison I, Buchdahl RM, Bush A. Primary ciliary dyskinesia: age at diagnosis and symptom history. Acta Paediatr. 2002;91:667–669. - PubMed
-
- Bush A, Hogg C, Mitchison HM, Nisbet M, Wilson R. Update in primary ciliary dyskinesia. Clin. Pulm. Med. 2009;16:219–225.
-
- Bush A. Congenital heart disease in primary ciliary dyskinesia. Pediatr. Cardiol. 1998;19:191. - PubMed
-
- Kennedy MP, et al. Congenital heart disease and other heterotaxic defects in a large cohort of patients with primary ciliary dyskinesia. Circulation. 2007;115:2814–2821. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

