A fatal mitochondrial disease is associated with defective NFU1 function in the maturation of a subset of mitochondrial Fe-S proteins
- PMID: 22077971
- PMCID: PMC3213398
- DOI: 10.1016/j.ajhg.2011.10.005
A fatal mitochondrial disease is associated with defective NFU1 function in the maturation of a subset of mitochondrial Fe-S proteins
Abstract
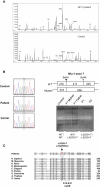
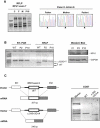
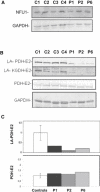
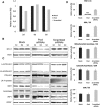
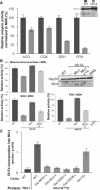
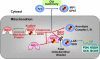
We report on ten individuals with a fatal infantile encephalopathy and/or pulmonary hypertension, leading to death before the age of 15 months. Hyperglycinemia and lactic acidosis were common findings. Glycine cleavage system and pyruvate dehydrogenase complex (PDHC) activities were low. Homozygosity mapping revealed a perfectly overlapping homozygous region of 1.24 Mb corresponding to chromosome 2 and led to the identification of a homozygous missense mutation (c.622G > T) in NFU1, which encodes a conserved protein suggested to participate in Fe-S cluster biogenesis. Nine individuals were homozygous for this mutation, whereas one was compound heterozygous for this and a splice-site (c.545 + 5G > A) mutation. The biochemical phenotype suggested an impaired activity of the Fe-S enzyme lipoic acid synthase (LAS). Direct measurement of protein-bound lipoic acid in individual tissues indeed showed marked decreases. Upon depletion of NFU1 by RNA interference in human cell culture, LAS and, in turn, PDHC activities were largely diminished. In addition, the amount of succinate dehydrogenase, but no other Fe-S proteins, was decreased. In contrast, depletion of the general Fe-S scaffold protein ISCU severely affected assembly of all tested Fe-S proteins, suggesting that NFU1 performs a specific function in mitochondrial Fe-S cluster maturation. Similar biochemical effects were observed in Saccharomyces cerevisiae upon deletion of NFU1, resulting in lower lipoylation and SDH activity. Importantly, yeast Nfu1 protein carrying the individuals' missense mutation was functionally impaired. We conclude that NFU1 functions as a late-acting maturation factor for a subset of mitochondrial Fe-S proteins.
Copyright © 2011 The American Society of Human Genetics. Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Assembly of the [4Fe-4S] cluster of NFU1 requires the coordinated donation of two [2Fe-2S] clusters from the scaffold proteins, ISCU2 and ISCA1.Hum Mol Genet. 2020 Nov 25;29(19):3165-3182. doi: 10.1093/hmg/ddaa172. Hum Mol Genet. 2020. PMID: 32776106 Free PMC article.
-
Protein expression profiles in patients carrying NFU1 mutations. Contribution to the pathophysiology of the disease.J Inherit Metab Dis. 2013 Sep;36(5):841-7. doi: 10.1007/s10545-012-9565-z. Epub 2012 Nov 22. J Inherit Metab Dis. 2013. PMID: 23179554
-
Impact of mutations within the [Fe-S] cluster or the lipoic acid biosynthesis pathways on mitochondrial protein expression profiles in fibroblasts from patients.Mol Genet Metab. 2017 Nov;122(3):85-94. doi: 10.1016/j.ymgme.2017.08.001. Epub 2017 Aug 3. Mol Genet Metab. 2017. PMID: 28803783
-
Mitochondrial iron-sulfur protein biogenesis and human disease.Biochimie. 2014 May;100:61-77. doi: 10.1016/j.biochi.2014.01.010. Epub 2014 Jan 23. Biochimie. 2014. PMID: 24462711 Review.
-
A hub for regulation of mitochondrial metabolism: Fatty acid and lipoic acid biosynthesis.IUBMB Life. 2024 Jun;76(6):332-344. doi: 10.1002/iub.2802. Epub 2023 Dec 13. IUBMB Life. 2024. PMID: 38088214 Review.
Cited by
-
Recent progress of methods for cuproptosis detection.Front Mol Biosci. 2024 Sep 4;11:1460987. doi: 10.3389/fmolb.2024.1460987. eCollection 2024. Front Mol Biosci. 2024. PMID: 39297074 Free PMC article. Review.
-
Mitochondrial Bol1 and Bol3 function as assembly factors for specific iron-sulfur proteins.Elife. 2016 Aug 17;5:e16673. doi: 10.7554/eLife.16673. Elife. 2016. PMID: 27532772 Free PMC article.
-
Assembly of the [4Fe-4S] cluster of NFU1 requires the coordinated donation of two [2Fe-2S] clusters from the scaffold proteins, ISCU2 and ISCA1.Hum Mol Genet. 2020 Nov 25;29(19):3165-3182. doi: 10.1093/hmg/ddaa172. Hum Mol Genet. 2020. PMID: 32776106 Free PMC article.
-
The iron-sulfur cluster assembly machineries in plants: current knowledge and open questions.Front Plant Sci. 2013 Jul 24;4:259. doi: 10.3389/fpls.2013.00259. eCollection 2013. Front Plant Sci. 2013. PMID: 23898337 Free PMC article.
-
The role of mitochondria in cellular iron-sulfur protein biogenesis: mechanisms, connected processes, and diseases.Cold Spring Harb Perspect Biol. 2013 Aug 1;5(8):a011312. doi: 10.1101/cshperspect.a011312. Cold Spring Harb Perspect Biol. 2013. PMID: 23906713 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

