Genetic landscape of high hyperdiploid childhood acute lymphoblastic leukemia
- PMID: 21098271
- PMCID: PMC3003126
- DOI: 10.1073/pnas.1006981107
Genetic landscape of high hyperdiploid childhood acute lymphoblastic leukemia
Abstract
High hyperdiploid acute lymphoblastic leukemia (ALL) is one of the most common malignancies in children. It is characterized by gain of chromosomes, typically +X, +4, +6, +10, +14, +17, +18, and +21,+21; little is known about additional genetic aberrations. Approximately 20% of the patients relapse; therefore it is clinically important to identify risk-stratifying markers. We used SNP array analysis to investigate a consecutive series of 74 cases of high hyperdiploid ALL. We show that the characteristic chromosomal gains are even more frequent than previously believed, indicating that karyotyping mistakes are common, and that almost 80% of the cases display additional abnormalities detectable by SNP array analysis. Subclonality analysis strongly implied that the numerical aberrations were primary and arose before structural events, suggesting that step-wise evolution of the leukemic clone is common. An association between duplication of 1q and +5 was seen (P = 0.003). Other frequent abnormalities included whole-chromosome uniparental isodisomies (wUPIDs) 9 and 11, gain of 17q not associated with isochromosome formation, extra gain of part of 21q, deletions of ETS variant 6 (ETV6), cyclin-dependent kinase inhibitor 2A (CKDN2A) and paired box 5 (PAX5), and PAN3 poly(A) specific ribonuclease subunit homolog (PAN3) microdeletions. Comparison of whole-chromosome and partial UPID9 suggested different pathogenetic outcomes, with the former not involving CDKN2A. Finally, two cases had partial deletions of AT rich interactive domain 5B (ARID5B), indicating that acquired as well as constitutional variants in this locus may be associated with pediatric ALL. Here we provide a comprehensive characterization of the genetic landscape of high hyperdiploid childhood ALL, including the heterogeneous pattern of secondary genetic events.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
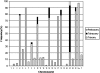


Similar articles
-
Heterogeneity of hyperdiploid (51-67) childhood acute lymphoblastic leukemia.Leukemia. 1996 Feb;10(2):213-24. Leukemia. 1996. PMID: 8637229
-
Identification of numerical and structural chromosome aberrations in 15 high hyperdiploid childhood acute lymphoblastic leukemias using spectral karyotyping.Eur J Haematol. 2001 May;66(5):297-304. doi: 10.1034/j.1600-0609.2001.066005297.x. Eur J Haematol. 2001. PMID: 11422408
-
Relapsed childhood high hyperdiploid acute lymphoblastic leukemia: presence of preleukemic ancestral clones and the secondary nature of microdeletions and RTK-RAS mutations.Leukemia. 2010 May;24(5):924-31. doi: 10.1038/leu.2010.39. Epub 2010 Mar 18. Leukemia. 2010. PMID: 20237506
-
High hyperdiploid childhood acute lymphoblastic leukemia.Genes Chromosomes Cancer. 2009 Aug;48(8):637-60. doi: 10.1002/gcc.20671. Genes Chromosomes Cancer. 2009. PMID: 19415723 Review.
-
Diagnostic and prognostic significance of chromosome abnormalities in childhood acute lymphoblastic leukemia.Ann N Y Acad Sci. 1997 Sep 17;824:8-27. doi: 10.1111/j.1749-6632.1997.tb46206.x. Ann N Y Acad Sci. 1997. PMID: 9382457 Review.
Cited by
-
ARID5B and IKZF1 variants, selected demographic factors, and childhood acute lymphoblastic leukemia: a report from the Children's Oncology Group.Leuk Res. 2013 Aug;37(8):936-42. doi: 10.1016/j.leukres.2013.04.022. Epub 2013 May 18. Leuk Res. 2013. PMID: 23692655 Free PMC article.
-
Defining low-risk high hyperdiploidy in patients with paediatric acute lymphoblastic leukaemia: a retrospective analysis of data from the UKALL97/99 and UKALL2003 clinical trials.Lancet Haematol. 2021 Nov;8(11):e828-e839. doi: 10.1016/S2352-3026(21)00304-5. Lancet Haematol. 2021. PMID: 34715050 Free PMC article. Clinical Trial.
-
Germline and somatic drivers in inherited hematologic malignancies.Front Oncol. 2023 Oct 13;13:1205855. doi: 10.3389/fonc.2023.1205855. eCollection 2023. Front Oncol. 2023. PMID: 37904876 Free PMC article. Review.
-
Genetic predisposition to B-cell acute lymphoblastic leukemia at 14q11.2 is mediated by a CEBPE promoter polymorphism.Leukemia. 2019 Jan;33(1):1-14. doi: 10.1038/s41375-018-0184-z. Epub 2018 Jul 6. Leukemia. 2019. PMID: 29977016 Free PMC article.
-
Genetic and regulatory mechanism of susceptibility to high-hyperdiploid acute lymphoblastic leukaemia at 10p21.2.Nat Commun. 2017 Mar 3;8:14616. doi: 10.1038/ncomms14616. Nat Commun. 2017. PMID: 28256501 Free PMC article.
References
-
- Hjalgrim LL, et al. Age- and sex-specific incidence of childhood leukemia by immunophenotype in the Nordic countries. J Natl Cancer Inst. 2003;95:1539–1544. - PubMed
-
- Paulsson K, Johansson B. High hyperdiploid childhood acute lymphoblastic leukemia. Genes Chromosomes Cancer. 2009;48:637–660. - PubMed
-
- Third International Workshop on Chromosomes in Leukemia Chromosomal abnormalities in acute lymphoblastic leukemia: Structural and numerical changes in 234 cases. Cancer Genet Cytogenet. 1981;4:101–110. - PubMed
-
- Groupe Francais de Cytogénétique Hématologique Collaborative study of karyotypes in childhood acute lymphoblastic leukemias. Leukemia. 1993;7:10–19. - PubMed
-
- Moorman AV, et al. United Kingdom Medical Research Council's Childhood Leukemia Working Party Outcome heterogeneity in childhood high-hyperdiploid acute lymphoblastic leukemia. Blood. 2003;102:2756–2762. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

