Mechanisms of change in gene copy number
- PMID: 19597530
- PMCID: PMC2864001
- DOI: 10.1038/nrg2593
Mechanisms of change in gene copy number
Abstract
Deletions and duplications of chromosomal segments (copy number variants, CNVs) are a major source of variation between individual humans and are an underlying factor in human evolution and in many diseases, including mental illness, developmental disorders and cancer. CNVs form at a faster rate than other types of mutation, and seem to do so by similar mechanisms in bacteria, yeast and humans. Here we review current models of the mechanisms that cause copy number variation. Non-homologous end-joining mechanisms are well known, but recent models focus on perturbation of DNA replication and replication of non-contiguous DNA segments. For example, cellular stress might induce repair of broken replication forks to switch from high-fidelity homologous recombination to non-homologous repair, thus promoting copy number change.
Figures
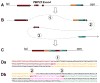
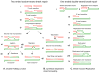
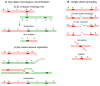


Similar articles
-
Replication stress induces genome-wide copy number changes in human cells that resemble polymorphic and pathogenic variants.Am J Hum Genet. 2009 Mar;84(3):339-50. doi: 10.1016/j.ajhg.2009.01.024. Epub 2009 Feb 19. Am J Hum Genet. 2009. PMID: 19232554 Free PMC article.
-
Analysis of copy number variants and segmental duplications in the human genome: Evidence for a change in the process of formation in recent evolutionary history.Genome Res. 2008 Dec;18(12):1865-74. doi: 10.1101/gr.081422.108. Epub 2008 Oct 8. Genome Res. 2008. PMID: 18842824 Free PMC article.
-
An Evolutionary Perspective on the Impact of Genomic Copy Number Variation on Human Health.J Mol Evol. 2020 Jan;88(1):104-119. doi: 10.1007/s00239-019-09911-6. Epub 2019 Sep 14. J Mol Evol. 2020. PMID: 31522275 Review.
-
Ionizing radiation and genetic risks XIV. Potential research directions in the post-genome era based on knowledge of repair of radiation-induced DNA double-strand breaks in mammalian somatic cells and the origin of deletions associated with human genomic disorders.Mutat Res. 2005 Oct 15;578(1-2):333-70. doi: 10.1016/j.mrfmmm.2005.06.020. Epub 2005 Aug 5. Mutat Res. 2005. PMID: 16084534 Review.
-
A microhomology-mediated break-induced replication model for the origin of human copy number variation.PLoS Genet. 2009 Jan;5(1):e1000327. doi: 10.1371/journal.pgen.1000327. Epub 2009 Jan 30. PLoS Genet. 2009. PMID: 19180184 Free PMC article. Review.
Cited by
-
A common copy-number breakpoint of ERBB2 amplification in breast cancer colocalizes with a complex block of segmental duplications.Breast Cancer Res. 2012 Nov 26;14(6):R150. doi: 10.1186/bcr3362. Breast Cancer Res. 2012. PMID: 23181561 Free PMC article.
-
R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli.Nat Commun. 2013;4:2115. doi: 10.1038/ncomms3115. Nat Commun. 2013. PMID: 23828459 Free PMC article.
-
Making Choices: DNA Replication Fork Recovery Mechanisms.Semin Cell Dev Biol. 2021 May;113:27-37. doi: 10.1016/j.semcdb.2020.10.001. Epub 2020 Oct 22. Semin Cell Dev Biol. 2021. PMID: 33967572 Free PMC article. Review.
-
PAV markers in Sorghum bicolour: genome pattern, affected genes and pathways, and genetic linkage map construction.Theor Appl Genet. 2015 Apr;128(4):623-37. doi: 10.1007/s00122-015-2458-4. Epub 2015 Jan 30. Theor Appl Genet. 2015. PMID: 25634103 Free PMC article.
-
Inevitability and containment of replication errors for eukaryotic genome lengths spanning megabase to gigabase.Proc Natl Acad Sci U S A. 2016 Sep 27;113(39):E5765-74. doi: 10.1073/pnas.1603241113. Epub 2016 Sep 14. Proc Natl Acad Sci U S A. 2016. PMID: 27630194 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
- R01 GM064022/GM/NIGMS NIH HHS/United States
- R01 GM80600/GM/NIGMS NIH HHS/United States
- R01 GM053158-14/GM/NIGMS NIH HHS/United States
- M01 RR000188/RR/NCRR NIH HHS/United States
- R01 GM064022-07/GM/NIGMS NIH HHS/United States
- R01 NS058529-01A2/NS/NINDS NIH HHS/United States
- R01 NS59529/NS/NINDS NIH HHS/United States
- R01 CA085777/CA/NCI NIH HHS/United States
- R01 GM053158-13/GM/NIGMS NIH HHS/United States
- R01 CA85777/CA/NCI NIH HHS/United States
- R01 GM53158/GM/NIGMS NIH HHS/United States
- R21 NS059529/NS/NINDS NIH HHS/United States
- R01 GM080600-03/GM/NIGMS NIH HHS/United States
- R01 GM080600/GM/NIGMS NIH HHS/United States
- R01 GM053158/GM/NIGMS NIH HHS/United States
- R01 CA085777-05/CA/NCI NIH HHS/United States
- R01 GM64022/GM/NIGMS NIH HHS/United States
- R01 NS058529/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

