The TREX1 double-stranded DNA degradation activity is defective in dominant mutations associated with autoimmune disease
- PMID: 18805785
- PMCID: PMC2581595
- DOI: 10.1074/jbc.M806155200
The TREX1 double-stranded DNA degradation activity is defective in dominant mutations associated with autoimmune disease
Abstract
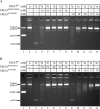
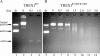
Mutations in TREX1 have been linked to a spectrum of human autoimmune diseases including Aicardi-Goutières syndrome (AGS), familial chilblain lupus (FCL), systemic lupus erythematosus, and retinal vasculopathy and cerebral leukodystrophy. A common feature in these conditions is the frequent detection of antibodies to double-stranded DNA (dsDNA). TREX1 participates in a cell death process implicating this major 3' --> 5' exonuclease in genomic DNA degradation to minimize potential immune activation by persistent self DNA. The TREX1 D200N and D18N dominant heterozygous mutations were identified in AGS and FCL, respectively. TREX1 enzymes containing the D200N and D18N mutations were compared using nicked dsDNA and single-stranded DNA (ssDNA) degradation assays. The TREX1WT/D200N and TREX1WT/D18N heterodimers are completely deficient at degrading dsDNA and degrade ssDNA at an expected approximately 2-fold lower rate than TREX1WT enzyme. Further, the D200N- and D18N-containing TREX1 homo- and heterodimers inhibit the dsDNA degradation activity of TREX1WT enzyme, providing a likely explanation for the dominant phenotype of these TREX1 mutant alleles in AGS and FCL. By comparison, the TREX1 R114H homozygous mutation causes AGS and is found as a heterozygous mutation in systemic lupus erythematosus. The TREX1R114H/R114H homodimer has dysfunctional dsDNA and ssDNA degradation activities and does not detectibly inhibit the TREX1WT enzyme, whereas the TREX1WT/R114H heterodimer has a functional dsDNA degradation activity, supporting the recessive genetics of TREX1 R114H in AGS. The dysfunctional dsDNA degradation activities of these disease-related TREX1 mutants could account for persistent dsDNA from dying cells leading to an aberrant immune response in these clinically related disorders.
Figures
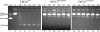

Similar articles
-
Defects in DNA degradation revealed in crystal structures of TREX1 exonuclease mutations linked to autoimmune disease.DNA Repair (Amst). 2012 Jan 2;11(1):65-73. doi: 10.1016/j.dnarep.2011.10.007. Epub 2011 Nov 8. DNA Repair (Amst). 2012. PMID: 22071149 Free PMC article.
-
The TREX1 exonuclease R114H mutation in Aicardi-Goutières syndrome and lupus reveals dimeric structure requirements for DNA degradation activity.J Biol Chem. 2011 Nov 18;286(46):40246-54. doi: 10.1074/jbc.M111.297903. Epub 2011 Sep 21. J Biol Chem. 2011. PMID: 21937424 Free PMC article.
-
The Arg-62 residues of the TREX1 exonuclease act across the dimer interface contributing to catalysis in the opposing protomers.J Biol Chem. 2014 Apr 18;289(16):11556-11565. doi: 10.1074/jbc.M114.559252. Epub 2014 Mar 9. J Biol Chem. 2014. PMID: 24616097 Free PMC article.
-
New roles for the major human 3'-5' exonuclease TREX1 in human disease.Cell Cycle. 2008 Jun 15;7(12):1718-25. doi: 10.4161/cc.7.12.6162. Epub 2008 Jun 16. Cell Cycle. 2008. PMID: 18583934 Free PMC article. Review.
-
TREX1 As a Potential Therapeutic Target for Autoimmune and Inflammatory Diseases.Curr Pharm Des. 2019;25(30):3239-3247. doi: 10.2174/1381612825666190902113218. Curr Pharm Des. 2019. PMID: 31475890 Review.
Cited by
-
TREX1 as a Novel Immunotherapeutic Target.Front Immunol. 2021 Apr 1;12:660184. doi: 10.3389/fimmu.2021.660184. eCollection 2021. Front Immunol. 2021. PMID: 33868310 Free PMC article.
-
Hepatitis B virus nucleocapsid uncoating: biological consequences and regulation by cellular nucleases.Emerg Microbes Infect. 2021 Dec;10(1):852-864. doi: 10.1080/22221751.2021.1919034. Emerg Microbes Infect. 2021. PMID: 33870849 Free PMC article.
-
Defects in DNA degradation revealed in crystal structures of TREX1 exonuclease mutations linked to autoimmune disease.DNA Repair (Amst). 2012 Jan 2;11(1):65-73. doi: 10.1016/j.dnarep.2011.10.007. Epub 2011 Nov 8. DNA Repair (Amst). 2012. PMID: 22071149 Free PMC article.
-
Recognition and processing of double-stranded DNA by ExoX, a distributive 3'-5' exonuclease.Nucleic Acids Res. 2013 Aug;41(15):7556-65. doi: 10.1093/nar/gkt495. Epub 2013 Jun 14. Nucleic Acids Res. 2013. PMID: 23771145 Free PMC article.
-
The landscape of rare genetic variants in familial Waldenström macroglobulinemia.Blood Neoplasia. 2024 Jun;1(2):100013. doi: 10.1016/j.bneo.2024.100013. Epub 2024 Apr 12. Blood Neoplasia. 2024. PMID: 39036705 Free PMC article.
References
-
- Mazur, D. J., and Perrino, F. W. (1999) J. Biol. Chem. 274 19655-19660 - PubMed
-
- Mazur, D. J., and Perrino, F. W. (2001) J. Biol. Chem. 276 17022-17029 - PubMed
-
- Mazur, D. J., and Perrino, F. W. (2001) J. Biol. Chem. 276 14718-14727 - PubMed
-
- Perrino, F. W., Miller, H., and Ealey, K. A. (1994) J. Biol. Chem. 269 16357-16363 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

