Human Tdp1 cleaves a broad spectrum of substrates, including phosphoamide linkages
- PMID: 16141202
- PMCID: PMC1351008
- DOI: 10.1074/jbc.M508898200
Human Tdp1 cleaves a broad spectrum of substrates, including phosphoamide linkages
Abstract
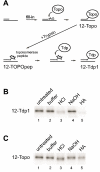
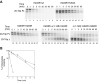
Human tyrosyl-DNA phosphodiesterase (Tdp1) hydrolyzes the phosphodiester bond between a DNA 3' end and a tyrosyl moiety. In eukaryotic cells, this type of linkage is found in stalled topoisomerase I-DNA covalent complexes, and Tdp1 has been implicated in the repair of such complexes in vivo. We confirm here that the Tdp1 catalytic cycle involves a covalent reaction intermediate in which a histidine residue is connected to a DNA 3'-phosphate through a phosphoamide linkage. Most surprisingly, this linkage can be hydrolyzed by Tdp1, and unlike a topoisomerase I-DNA complex, which requires modification to be an efficient substrate for Tdp1, the native form of Tdp1 can be removed from the DNA. The spinocerebellar ataxia with axonal neuropathy neurodegenerative disease is caused by the H493R mutant form of Tdp1, which shows reduced enzymatic activity and accumulates the Tdp1-DNA covalent intermediate. The ability of wild type Tdp1 to remove the stalled mutant protein from the DNA likely explains the recessive nature of spinocerebellar ataxia with axonal neuropathy. In addition to its activity on phosphotyrosine and phosphohistidine substrates, Tdp1 also possesses a limited DNA and RNA 3'-exonuclease activity in which a single nucleoside is removed from the 3'-hydroxyl end of the substrate. Furthermore, Tdp1 also removes a 3' abasic site and an artificial 3'-biotin adduct from the DNA. In combination with earlier data showing that Tdp1 can use 3'-phosphoglycolate as a substrate, these data suggest that Tdp1 may function to remove a variety of 3' adducts from DNA during DNA repair.
Figures
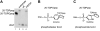



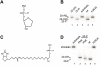
Similar articles
-
Tyrosyl-DNA phosphodiesterase (Tdp1) (3'-phosphotyrosyl DNA phosphodiesterase).Methods Enzymol. 2006;409:511-24. doi: 10.1016/S0076-6879(05)09030-0. Methods Enzymol. 2006. PMID: 16793421
-
In vitro complementation of Tdp1 deficiency indicates a stabilized enzyme-DNA adduct from tyrosyl but not glycolate lesions as a consequence of the SCAN1 mutation.DNA Repair (Amst). 2009 May 1;8(5):654-63. doi: 10.1016/j.dnarep.2008.12.012. Epub 2009 Feb 10. DNA Repair (Amst). 2009. PMID: 19211312 Free PMC article.
-
Tyrosyl-DNA phosphodiesterase I catalytic mutants reveal an alternative nucleophile that can catalyze substrate cleavage.J Biol Chem. 2015 Mar 6;290(10):6203-14. doi: 10.1074/jbc.M114.635284. Epub 2015 Jan 21. J Biol Chem. 2015. PMID: 25609251 Free PMC article.
-
Tyrosyl-DNA phosphodiesterase I resolves both naturally and chemically induced DNA adducts and its potential as a therapeutic target.Drug Metab Rev. 2014 Nov;46(4):494-507. doi: 10.3109/03602532.2014.971957. Epub 2014 Oct 20. Drug Metab Rev. 2014. PMID: 25327705 Review.
-
Tyrosyl-DNA phosphodiesterase as a target for anticancer therapy.Anticancer Agents Med Chem. 2008 May;8(4):381-9. doi: 10.2174/187152008784220357. Anticancer Agents Med Chem. 2008. PMID: 18473723 Free PMC article. Review.
Cited by
-
Structure-function studies of a plant tyrosyl-DNA phosphodiesterase provide novel insights into DNA repair mechanisms of Arabidopsis thaliana.Biochem J. 2012 Apr 1;443(1):49-56. doi: 10.1042/BJ20111308. Biochem J. 2012. PMID: 22214184 Free PMC article.
-
Synthesis and Biological Activities of 11- and 12-Substituted Benzophenanthridinone Derivatives as DNA Topoisomerase IB and Tyrosyl-DNA Phosphodiesterase 1 Inhibitors.ChemMedChem. 2023 May 16;18(10):e202200593. doi: 10.1002/cmdc.202200593. Epub 2023 Mar 17. ChemMedChem. 2023. PMID: 36932053 Free PMC article.
-
Tyrosyl-DNA Phosphodiesterase 1 (Tdp1) inhibitors.Expert Opin Ther Pat. 2011 Sep;21(9):1285-92. doi: 10.1517/13543776.2011.604314. Expert Opin Ther Pat. 2011. PMID: 21843105 Free PMC article. Review.
-
Identification of novel PARP inhibitors using a cell-based TDP1 inhibitory assay in a quantitative high-throughput screening platform.DNA Repair (Amst). 2014 Sep;21:177-82. doi: 10.1016/j.dnarep.2014.03.006. Epub 2014 Apr 29. DNA Repair (Amst). 2014. PMID: 24794403 Free PMC article.
-
Recognition and repair of chemically heterogeneous structures at DNA ends.Environ Mol Mutagen. 2015 Jan;56(1):1-21. doi: 10.1002/em.21892. Epub 2014 Aug 11. Environ Mol Mutagen. 2015. PMID: 25111769 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

