Mutations in the small GTP-ase late endosomal protein RAB7 cause Charcot-Marie-Tooth type 2B neuropathy
- PMID: 12545426
- PMCID: PMC1180247
- DOI: 10.1086/367847
Mutations in the small GTP-ase late endosomal protein RAB7 cause Charcot-Marie-Tooth type 2B neuropathy
Abstract
Charcot-Marie-Tooth type 2B (CMT2B) is clinically characterized by marked distal muscle weakness and wasting and a high frequency of foot ulcers, infections, and amputations of the toes because of recurrent infections. CMT2B maps to chromosome 3q13-q22. We refined the CMT2B locus to a 2.5-cM region and report two missense mutations (Leu129Phe and Val162Met) in the small GTP-ase late endosomal protein RAB7 which causes the CMT2B phenotype in three extended families and in three patients with a positive family history. The alignment of RAB7 orthologs shows that both missense mutations target highly conserved amino acid residues. RAB7 is ubiquitously expressed, and we found expression in sensory and motor neurons.
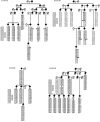
Figures


Similar articles
-
A novel RAB7 mutation in a Chinese family with Charcot-Marie-Tooth type 2B disease.Gene. 2014 Jan 25;534(2):431-4. Gene. 2014. PMID: 24498653
-
Rab7 and the CMT2B disease.Biochem Soc Trans. 2009 Oct;37(Pt 5):1027-31. doi: 10.1042/BST0371027. Biochem Soc Trans. 2009. PMID: 19754445 Review.
-
Functional characterization of Rab7 mutant proteins associated with Charcot-Marie-Tooth type 2B disease.J Neurosci. 2008 Feb 13;28(7):1640-8. doi: 10.1523/JNEUROSCI.3677-07.2008. J Neurosci. 2008. PMID: 18272684 Free PMC article.
-
Molecular basis of Charcot-Marie-Tooth type 2B disease.Biochem Soc Trans. 2012 Dec 1;40(6):1368-72. doi: 10.1042/BST20120197. Biochem Soc Trans. 2012. PMID: 23176482
-
Charcot Marie Tooth 2B Peripheral Sensory Neuropathy: How Rab7 Mutations Impact NGF Signaling?Int J Mol Sci. 2017 Feb 4;18(2):324. doi: 10.3390/ijms18020324. Int J Mol Sci. 2017. PMID: 28165391 Free PMC article. Review.
Cited by
-
Charcot-Marie-Tooth disease and intracellular traffic.Prog Neurobiol. 2012 Dec;99(3):191-225. doi: 10.1016/j.pneurobio.2012.03.003. Epub 2012 Mar 22. Prog Neurobiol. 2012. PMID: 22465036 Free PMC article. Review.
-
LRRK2-Related Parkinson's Disease Due to Altered Endolysosomal Biology With Variable Lewy Body Pathology: A Hypothesis.Front Neurosci. 2020 Jun 4;14:556. doi: 10.3389/fnins.2020.00556. eCollection 2020. Front Neurosci. 2020. PMID: 32581693 Free PMC article. Review.
-
Autophagic and endo-lysosomal dysfunction in neurodegenerative disease.Mol Brain. 2019 Nov 29;12(1):100. doi: 10.1186/s13041-019-0504-x. Mol Brain. 2019. PMID: 31783880 Free PMC article. Review.
-
Rab GTPases: Switching to Human Diseases.Cells. 2019 Aug 16;8(8):909. doi: 10.3390/cells8080909. Cells. 2019. PMID: 31426400 Free PMC article. Review.
-
Systematic discovery of Rab GTPases with synaptic functions in Drosophila.Curr Biol. 2011 Oct 25;21(20):1704-15. doi: 10.1016/j.cub.2011.08.058. Epub 2011 Oct 13. Curr Biol. 2011. PMID: 22000105 Free PMC article.
References
Electronic-Database Information
-
- ClustalW, http://npsa-pbil.ibcp.fr/ (for multiple protein alignment)
-
- GenBank, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=Protein (for LotusLink and protein sequences: RAB7_human [P51149]; Rab7_mouse [P51150]; Rab7_rat [P09527]; Rab-protein 7 Drosophila melanogaster [NP_524472]; Rab7_Dictyostelium discoideum [P36411]; Ras related protein_Caenorhabditis elegans [NP_496549]; Rab7_Arabidopsis thaliana [O04157]; and YPT7_YEAST [P32939])
-
- German Human Genome Project, http://www.rzpd.de/ (clones for partial human RAB7 cDNA sequences)
-
- NCBI Map Viewer, http://www.ncbi.nlm.nih.gov/mapview/map_search.cgi?chr=hum_chr.inf&query (for finding known genes, ESTs, and putative novel genes in the CMT2B region)
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for CMT2B [MIM 600882] and HSN I [MIM 162400]).
References
-
- Auer-Grumbach M, De Jonghe P, Verhoeven K, Timmerman V, Wagner K, Hartung HP, Nicholson GA. Autosomal dominant inherited neuropathies with prominent sensory loss and mutilations: a review. Arch Neurol (in press) - PubMed
-
- Auer-Grumbach M, De Jonghe P, Wagner K, Verhoeven K, Hartung HP, Timmerman V (2000a) Phenotype-genotype correlations in a CMT2B family with refined 3q13-q22 locus. Neurology 55:1552–1557 - PubMed
-
- Auer-Grumbach M, Wagner K, Timmerman V, De Jonghe P, Hartung HP (2000b) Ulcero-mutilating neuropathy in an Austrian kinship without linkage to hereditary motor and sensory neuropathy IIB and hereditary sensory neuropathy I loci. Neurology 54:45–52 - PubMed
-
- Bejaoui K, Wu C, Scheffler MD, Haan G, Ashby P, Wu L, de Jong P, Brown RH Jr (2001) SPTLC1 is mutated in hereditary sensory neuropathy, type 1. Nat Genet 27:261–262 - PubMed
-
- Bellone E, Rodolico C, Toscano A, Di Maria E, Cassandrini D, Pizzuti A, Pigullo S, Mazzeo A, Macaione V, Girlanda P, Vita G, Ajmar F, Mandich P (2002) A family with autosomal dominant mutilating neuropathy not linked to either Charcot-Marie-Tooth disease type 2B (CMT2B) or hereditary sensory neuropathy type I (HSN I) loci. Neuromuscul Disord 12:286–291 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

