Structure and evolution of the Smith-Magenis syndrome repeat gene clusters, SMS-REPs
- PMID: 11997339
- PMCID: PMC186597
- DOI: 10.1101/gr.82802
Structure and evolution of the Smith-Magenis syndrome repeat gene clusters, SMS-REPs
Abstract
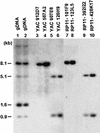
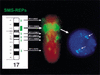
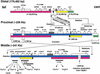
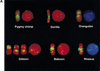
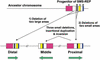
An approximately 4-Mb genomic segment on chromosome 17p11.2, commonly deleted in patients with the Smith-Magenis syndrome (SMS) and duplicated in patients with dup(17)(p11.2p11.2) syndrome, is flanked by large, complex low-copy repeats (LCRs), termed proximal and distal SMS-REP. A third copy, the middle SMS-REP, is located between them. SMS-REPs are believed to mediate nonallelic homologous recombination, resulting in both SMS deletions and reciprocal duplications. To delineate the genomic structure and evolutionary origin of SMS-REPs, we constructed a bacterial artificial chromosome/P1 artificial chromosome contig spanning the entire SMS region, including the SMS-REPs, determined its genomic sequence, and used fluorescence in situ hybridization to study the evolution of SMS-REP in several primate species. Our analysis shows that both the proximal SMS-REP (approximately 256 kb) and the distal copy (approximately 176 kb) are located in the same orientation and derived from a progenitor copy, whereas the middle SMS-REP (approximately 241 kb) is inverted and appears to have been derived from the proximal copy. The SMS-REP LCRs are highly homologous (>98%) and contain at least 14 genes/pseudogenes each. SMS-REPs are not present in mice and were duplicated after the divergence of New World monkeys from pre-monkeys approximately 40-65 million years ago. Our findings potentially explain why the vast majority of SMS deletions and dup(17)(p11.2p11.2) occur at proximal and distal SMS-REPs and further support previous observations that higher-order genomic architecture involving LCRs arose recently during primate speciation and may predispose the human genome to both meiotic and mitotic rearrangements.
Figures
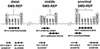
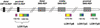

Similar articles
-
Reciprocal crossovers and a positional preference for strand exchange in recombination events resulting in deletion or duplication of chromosome 17p11.2.Am J Hum Genet. 2003 Dec;73(6):1302-15. doi: 10.1086/379979. Epub 2003 Nov 24. Am J Hum Genet. 2003. PMID: 14639526 Free PMC article.
-
Genes in a refined Smith-Magenis syndrome critical deletion interval on chromosome 17p11.2 and the syntenic region of the mouse.Genome Res. 2002 May;12(5):713-28. doi: 10.1101/gr.73702. Genome Res. 2002. PMID: 11997338 Free PMC article.
-
Genetic proof of unequal meiotic crossovers in reciprocal deletion and duplication of 17p11.2.Am J Hum Genet. 2002 Nov;71(5):1072-81. doi: 10.1086/344346. Epub 2002 Oct 9. Am J Hum Genet. 2002. PMID: 12375235 Free PMC article.
-
Trisomy 17p10-p12 due to mosaic supernumerary marker chromosome: delineation of molecular breakpoints and clinical phenotype, and comparison to other proximal 17p segmental duplications.Am J Med Genet A. 2005 Oct 1;138A(2):175-80. doi: 10.1002/ajmg.a.30948. Am J Med Genet A. 2005. PMID: 16152635 Review.
-
New developments in Smith-Magenis syndrome (del 17p11.2).Curr Opin Neurol. 2007 Apr;20(2):125-34. doi: 10.1097/WCO.0b013e3280895dba. Curr Opin Neurol. 2007. PMID: 17351481 Review.
Cited by
-
Modeling del(17)(p11.2p11.2) and dup(17)(p11.2p11.2) contiguous gene syndromes by chromosome engineering in mice: phenotypic consequences of gene dosage imbalance.Mol Cell Biol. 2003 May;23(10):3646-55. doi: 10.1128/MCB.23.10.3646-3655.2003. Mol Cell Biol. 2003. PMID: 12724422 Free PMC article.
-
Reciprocal crossovers and a positional preference for strand exchange in recombination events resulting in deletion or duplication of chromosome 17p11.2.Am J Hum Genet. 2003 Dec;73(6):1302-15. doi: 10.1086/379979. Epub 2003 Nov 24. Am J Hum Genet. 2003. PMID: 14639526 Free PMC article.
-
The breakpoint region of the most common isochromosome, i(17q), in human neoplasia is characterized by a complex genomic architecture with large, palindromic, low-copy repeats.Am J Hum Genet. 2004 Jan;74(1):1-10. doi: 10.1086/380648. Epub 2003 Dec 8. Am J Hum Genet. 2004. PMID: 14666446 Free PMC article.
-
Human BCAS3 expression in embryonic stem cells and vascular precursors suggests a role in human embryogenesis and tumor angiogenesis.PLoS One. 2007 Nov 21;2(11):e1202. doi: 10.1371/journal.pone.0001202. PLoS One. 2007. PMID: 18030336 Free PMC article.
-
Genes in a refined Smith-Magenis syndrome critical deletion interval on chromosome 17p11.2 and the syntenic region of the mouse.Genome Res. 2002 May;12(5):713-28. doi: 10.1101/gr.73702. Genome Res. 2002. PMID: 11997338 Free PMC article.
References
-
- Boerkoel CF, Inoue K, Reiter LT, Warner LE, Lupski JR. Molecular mechanisms for CMT1A duplication and HNPP deletion. Ann NY Acad Sci. 1999;883:22–35. - PubMed
-
- Cai WW, Reneker J, Chow C-W, Vaishnav M, Bradley A. An anchored framework BAC map of mouse chromosome 11 assembled using multiplex oligonucleotide hybridization. Genomics. 1998;54:387–397. - PubMed
-
- Chen K-S, Manian P, Koeuth T, Potocki L, Zhao Q, Chinault AC, Lee CC, Lupski JR. Homologous recombination of a flanking repeat gene cluster is a mechanism for a common contiguous gene deletion syndrome. Nat Genet. 1997;17:154–163. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
