In vitro 3'-end endonucleolytic processing defect in a human mitochondrial tRNA(Ser(UCN)) precursor with the U7445C substitution, which causes non-syndromic deafness
- PMID: 11691920
- PMCID: PMC60182
- DOI: 10.1093/nar/29.21.4334
In vitro 3'-end endonucleolytic processing defect in a human mitochondrial tRNA(Ser(UCN)) precursor with the U7445C substitution, which causes non-syndromic deafness
Abstract
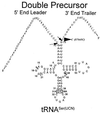
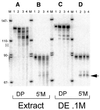
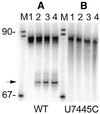
Eukaryotic tRNAs are transcribed as precursors. A 5'-end leader and 3'-end trailer are endonucleolytically removed by RNase P and 3'-tRNase before 3'-end CCA addition, aminoacylation, nuclear export and translation. 3'-End -CC can be a 3'-tRNase anti-determinant with the ability to prevent mature tRNA from recycling through 3'-tRNase. Twenty-two tRNAs punctuate the two rRNAs and 13 mRNAs in long, bidirectional mitochondrial transcripts. Accurate mitochondrial gene expression thus depends on endonucleolytic excision of tRNAs. Various mitochondrial diseases and syndromes could arise from defective tRNA end processing. The U7445C substitution in the human mitochondrial L-strand transcript (U74C directly following the discriminator base of tRNA(Ser(UCN))) causes non-syndromic deafness. The sequence of the precursor (G/UCU) becomes G/CCU, resembling a 3'-tRNase anti-determinant. We demonstrate that a tRNA(Ser(UCN)) precursor with the U7445C substitution cannot be processed in vitro by 3'-tRNase from human mitochondria. A 3'-end processing defect in this tRNA precursor could thus be responsible for mitochondrial disease.
Figures
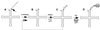
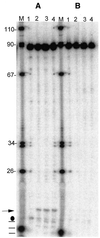

Similar articles
-
Mitochondrial tRNA 3' end metabolism and human disease.Nucleic Acids Res. 2004 Oct 11;32(18):5430-41. doi: 10.1093/nar/gkh884. Print 2004. Nucleic Acids Res. 2004. PMID: 15477393 Free PMC article. Review.
-
Naturally occurring mutations in human mitochondrial pre-tRNASer(UCN) can affect the transfer ribonuclease Z cleavage site, processing kinetics, and substrate secondary structure.J Biol Chem. 2006 Feb 17;281(7):3926-35. doi: 10.1074/jbc.M509822200. Epub 2005 Dec 16. J Biol Chem. 2006. PMID: 16361254
-
Pathogenesis-related mutations in the T-loops of human mitochondrial tRNAs affect 3' end processing and tRNA structure.RNA Biol. 2012 Mar;9(3):283-91. doi: 10.4161/rna.19025. Epub 2012 Mar 1. RNA Biol. 2012. PMID: 22336717 Free PMC article.
-
A new mechanism for mtDNA pathogenesis: impairment of post-transcriptional maturation leads to severe depletion of mitochondrial tRNASer(UCN) caused by T7512C and G7497A point mutations.Nucleic Acids Res. 2005 Sep 30;33(17):5647-58. doi: 10.1093/nar/gki876. Print 2005. Nucleic Acids Res. 2005. PMID: 16199753 Free PMC article.
-
[Mutations of mitochondrial tRNASer(UCN) and their connection with hearing loss].Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2017 Feb 10;34(1):128-132. doi: 10.3760/cma.j.issn.1003-9406.2017.01.030. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2017. PMID: 28186612 Review. Chinese.
Cited by
-
Gene rearrangements and evolution of tRNA pseudogenes in the mitochondrial genome of the parrotfish (Teleostei: Perciformes: Scaridae).J Mol Evol. 2004 Sep;59(3):287-97. doi: 10.1007/s00239-004-2621-z. J Mol Evol. 2004. PMID: 15553084
-
Failures in mitochondrial tRNAMet and tRNAGln metabolism caused by the novel 4401A>G mutation are involved in essential hypertension in a Han Chinese Family.Hypertension. 2009 Aug;54(2):329-37. doi: 10.1161/HYPERTENSIONAHA.109.129270. Epub 2009 Jun 22. Hypertension. 2009. PMID: 19546379 Free PMC article.
-
Optimization and characterization of tRNA-shRNA expression constructs.Nucleic Acids Res. 2007;35(8):2620-8. doi: 10.1093/nar/gkm103. Epub 2007 Apr 10. Nucleic Acids Res. 2007. PMID: 17426139 Free PMC article.
-
Transfer RNA detection by small RNA deep sequencing and disease association with myelodysplastic syndromes.BMC Genomics. 2015 Sep 24;16:727. doi: 10.1186/s12864-015-1929-y. BMC Genomics. 2015. PMID: 26400237 Free PMC article.
-
Mitochondrial tRNA 3' end metabolism and human disease.Nucleic Acids Res. 2004 Oct 11;32(18):5430-41. doi: 10.1093/nar/gkh884. Print 2004. Nucleic Acids Res. 2004. PMID: 15477393 Free PMC article. Review.
References
-
- Guerrier-Takada C., Gardiner,K., Marsh,T., Pace,N. and Altman,S. (1983) The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell, 35, 849–857. - PubMed
-
- Castaño J.G., Tobian,J.A. and Zasloff,M. (1985) Purification and characterization of an endonuclease from Xenopus laevis ovaries which accurately processes the 3′ terminus of human pre-tRNAi–Met (3′Pre-tRNase). J. Biol. Chem., 260, 9002–9008. - PubMed
-
- Frendewey D., Dingermann,T., Cooley,L. and Söll,D. (1985) Processing of precursor tRNAs in Drosophila. Processing of the 3′ end involves an endonucleolytic cleavage and occurs after 5′ end maturation. J. Biol. Chem., 260, 449–454. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

