Haploinsufficiency for one COL3A1 allele of type III procollagen results in a phenotype similar to the vascular form of Ehlers-Danlos syndrome, Ehlers-Danlos syndrome type IV
- PMID: 11577371
- PMCID: PMC1274375
- DOI: 10.1086/324123
Haploinsufficiency for one COL3A1 allele of type III procollagen results in a phenotype similar to the vascular form of Ehlers-Danlos syndrome, Ehlers-Danlos syndrome type IV
Abstract
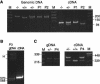
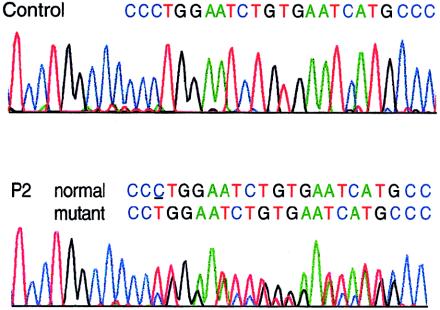
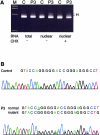
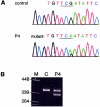
Mutations in the COL3A1 gene that encodes the chains of type III procollagen result in the vascular form of Ehlers-Danlos syndrome (EDS), EDS type IV, if they alter the sequence in the triple-helical domain. Although other fibrillar collagen-gene mutations that lead to allele instability or failure to incorporate proalpha-chains into trimers-and that thus reduce the amount of mature molecules produced-result in clinically apparent phenotypes, no such mutations have been identified in COL3A1. Furthermore, mice heterozygous for Col3a1 "null" alleles have no identified phenotype. We have now found three frameshift mutations (1832delAA, 413delC, and 555delT) that lead to premature termination codons (PTCs) in exons 27, 6, and 9, respectively, and to allele-product instability. The mRNA from each mutant allele was transcribed efficiently but rapidly degraded, presumably by the mechanisms of nonsense-mediated decay. In a fourth patient, we identified a point mutation, in the final exon, that resulted in a PTC (4294C-->T [Arg1432Ter]). In this last instance, the mRNA was stable but led to synthesis of a truncated protein that was not incorporated into mature type III procollagen molecules. In all probands, the presenting feature was vascular aneurysm or rupture. Thus, in contrast to mutations in genes that encode the dominant protein of a tissue (e.g., COL1A1 and COL2A1), in which "null" mutations result in phenotypes milder than those caused by mutations that alter protein sequence, the phenotypes produced by these mutations in COL3A1 overlap with those of the vascular form of EDS. This suggests that the major effect of many of these dominant mutations in the "minor" collagen genes may be expressed through protein deficiency rather than through incorporation of structurally altered molecules into fibrils.
Figures
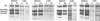
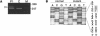
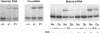
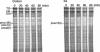
Similar articles
-
Abnormal type III collagen produced by an exon-17-skipping mutation of the COL3A1 gene in Ehlers-Danlos syndrome type IV is not incorporated into the extracellular matrix.Biochem J. 1995 Nov 1;311 ( Pt 3)(Pt 3):939-43. doi: 10.1042/bj3110939. Biochem J. 1995. PMID: 7487954 Free PMC article.
-
A base substitution at a splice site in the COL3A1 gene causes exon skipping and generates abnormal type III procollagen in a patient with Ehlers-Danlos syndrome type IV.J Biol Chem. 1990 Oct 5;265(28):17070-7. J Biol Chem. 1990. PMID: 2145268
-
Null alleles of the COL5A1 gene of type V collagen are a cause of the classical forms of Ehlers-Danlos syndrome (types I and II).Am J Hum Genet. 2000 Jun;66(6):1757-65. doi: 10.1086/302933. Epub 2000 May 4. Am J Hum Genet. 2000. PMID: 10796876 Free PMC article.
-
Type III collagen (COL3A1): Gene and protein structure, tissue distribution, and associated diseases.Gene. 2019 Jul 30;707:151-171. doi: 10.1016/j.gene.2019.05.003. Epub 2019 May 7. Gene. 2019. PMID: 31075413 Free PMC article. Review.
-
Inherited disorders of collagen gene structure and expression.Am J Med Genet. 1989 Sep;34(1):72-80. doi: 10.1002/ajmg.1320340114. Am J Med Genet. 1989. PMID: 2683783 Review.
Cited by
-
Gene panel sequencing in heritable thoracic aortic disorders and related entities - results of comprehensive testing in a cohort of 264 patients.Orphanet J Rare Dis. 2015 Feb 3;10:9. doi: 10.1186/s13023-014-0221-6. Orphanet J Rare Dis. 2015. PMID: 25644172 Free PMC article.
-
Vascular Ehlers-Danlos syndrome mutations in type III collagen differently stall the triple helical folding.J Biol Chem. 2013 Jun 28;288(26):19166-76. doi: 10.1074/jbc.M113.462002. Epub 2013 May 3. J Biol Chem. 2013. PMID: 23645670 Free PMC article.
-
GPR56 and the developing cerebral cortex: cells, matrix, and neuronal migration.Mol Neurobiol. 2013 Feb;47(1):186-96. doi: 10.1007/s12035-012-8343-0. Epub 2012 Sep 22. Mol Neurobiol. 2013. PMID: 23001883 Free PMC article. Review.
-
Identification of COL4A4 variants in Chinese patients with familial hematuria.Front Genet. 2023 Jan 9;13:1064491. doi: 10.3389/fgene.2022.1064491. eCollection 2022. Front Genet. 2023. PMID: 36699462 Free PMC article.
-
Biomarkers in EndoVascular Aneurysm Repair (EVAR) and Abdominal Aortic Aneurysm: Pathophysiology and Clinical Implications.Diagnostics (Basel). 2022 Jan 13;12(1):183. doi: 10.3390/diagnostics12010183. Diagnostics (Basel). 2022. PMID: 35054350 Free PMC article. Review.
References
Electronic-Database Information
-
- GenBank, http://www.ncbi.nlm.nih.gov/Genbank/ (for cDNA reference sequence [accession number NM_000090])
-
- Human Genome Project Working Draft at UCSC, http://genome.ucsc.edu/ (for COL3A1 gene sequence, chromosome 2 [accession numbers 193023938–193062278])
-
- Mutations in COL3A1,http://www.le.ac.uk/genetics/collagen/col3a1.html
-
- Online Mendelian Inheritance in Man (OMIM), http://www3.ncbi.nlm.nih.gov/Omim/ (for EDS type IV [MIM 130050])
References
-
- Ala-Kokko L, Kontusaari S, Baldwin CT, Kuivaniemi H, Prockop DJ (1989) Structure of cDNA clones coding for the entire prepro alpha 1 (III) chain of human type III procollagen: differences in protein structure from type I procollagen and conservation of codon preferences. Biochem J 260:509–516 - PMC - PubMed
-
- Beighton P, De Paepe A, Steinmann B, Tsipouras P, Wenstrup RJ (1998) Ehlers-Danlos syndromes: revised nosology, Villefranche, 1997: Ehlers-Danlos National Foundation (USA) and Ehlers-Danlos Support Group (UK). Am J Med Genet 77:31–37 - PubMed
-
- Benson-Chanda V, Su MW, Weil D, Chu ML, Ramirez F (1989) Cloning and analysis of the 5′ portion of the human type-III procollagen gene (COL3A1). Gene 78:255–265 - PubMed
-
- Bonadio J, Holbrook KA, Gelinas RE, Jacob J, Byers PH (1985) Altered triple helical structure of type I procollagen in lethal perinatal osteogenesis imperfecta. J Biol Chem 260:1734–1742 - PubMed
-
- Byers PH, Schwarze U. Ehlers-Danlos syndrome. In: Rimoin DL, Connor JM, Pyeritz RE (eds) Emery and Rimoin's principles and practice of medical genetics. Harcourt, London (in press)
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

