Human and mouse homologs of Schizosaccharomyces pombe rad1(+) and Saccharomyces cerevisiae RAD17: linkage to checkpoint control and mammalian meiosis
- PMID: 9716408
- PMCID: PMC317084
- DOI: 10.1101/gad.12.16.2560
Human and mouse homologs of Schizosaccharomyces pombe rad1(+) and Saccharomyces cerevisiae RAD17: linkage to checkpoint control and mammalian meiosis
Abstract
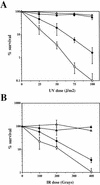
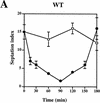
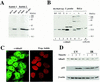
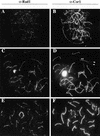
Preventing or delaying progress through the cell cycle in response to DNA damage is crucial for eukaryotic cells to allow the damage to be repaired and not incorporated irrevocably into daughter cells. Several genes involved in this process have been discovered in fission and budding yeast. Here, we report the identification of human and mouse homologs of the Schizosaccharomyces pombe DNA damage checkpoint control gene rad1(+) and its Saccharomyces cerevisiae homolog RAD17. The human gene HRAD1 is located on chromosome 5p13 and is most homologous to S. pombe rad1(+). This gene encodes a 382-amino-acid residue protein that is localized mainly in the nucleus and is expressed at high levels in proliferative tissues. This human gene significantly complements the sensitivity to UV light of a S. pombe strain mutated in rad1(+). Moreover, HRAD1 complements the checkpoint control defect of this strain after UV exposure. In addition to functioning in DNA repair checkpoints, S. cerevisiae RAD17 plays a role during meiosis to prevent progress through prophase I when recombination is interrupted. Consistent with a similar role in mammals, Rad1 protein is abundant in testis, and is associated with both synapsed and unsynapsed chromosomes during meiotic prophase I of spermatogenesis, with a staining pattern distinct from that of the recombination proteins Rad51 and Dmc1. Together, these data imply an important role for hRad1 both in the mitotic DNA damage checkpoint and in meiotic checkpoint mechanisms, and suggest that these events are highly conserved from yeast to humans.
Figures









Similar articles
-
A human and mouse homolog of the Schizosaccharomyces pombe rad1+ cell cycle checkpoint control gene.Genomics. 1998 Dec 1;54(2):331-7. doi: 10.1006/geno.1998.5582. Genomics. 1998. PMID: 9828137
-
RAD1, a human structural homolog of the Schizosaccharomyces pombe RAD1 cell cycle checkpoint gene.Genomics. 1998 Dec 1;54(2):344-7. doi: 10.1006/geno.1998.5589. Genomics. 1998. PMID: 9828139
-
HRAD1 and MRAD1 encode mammalian homologues of the fission yeast rad1(+) cell cycle checkpoint control gene.Nucleic Acids Res. 1998 Sep 1;26(17):3971-6. doi: 10.1093/nar/26.17.3971. Nucleic Acids Res. 1998. PMID: 9705507 Free PMC article.
-
Dial 9-1-1 for DNA damage: the Rad9-Hus1-Rad1 (9-1-1) clamp complex.DNA Repair (Amst). 2004 Aug-Sep;3(8-9):1009-14. doi: 10.1016/j.dnarep.2004.03.032. DNA Repair (Amst). 2004. PMID: 15279787 Review.
-
Repair of UV damage in the fission yeast Schizosaccharomyces pombe.Mutat Res. 2000 Jun 30;451(1-2):197-210. doi: 10.1016/s0027-5107(00)00050-6. Mutat Res. 2000. PMID: 10915873 Review.
Cited by
-
p53 binding protein 1 (53BP1) is an early participant in the cellular response to DNA double-strand breaks.J Cell Biol. 2000 Dec 25;151(7):1381-90. doi: 10.1083/jcb.151.7.1381. J Cell Biol. 2000. PMID: 11134068 Free PMC article.
-
Response of Xenopus Cds1 in cell-free extracts to DNA templates with double-stranded ends.Mol Biol Cell. 2000 May;11(5):1535-46. doi: 10.1091/mbc.11.5.1535. Mol Biol Cell. 2000. PMID: 10793133 Free PMC article.
-
Targeted deletion of mouse Rad1 leads to deficient cellular DNA damage responses.Protein Cell. 2011 May;2(5):410-22. doi: 10.1007/s13238-011-1049-7. Epub 2011 Jun 2. Protein Cell. 2011. PMID: 21637962 Free PMC article.
-
Meiotic DNA replication checkpoint control in fission yeast.Genes Dev. 1999 Oct 1;13(19):2581-93. doi: 10.1101/gad.13.19.2581. Genes Dev. 1999. PMID: 10521402 Free PMC article.
-
Inactivation of mouse Hus1 results in genomic instability and impaired responses to genotoxic stress.Genes Dev. 2000 Aug 1;14(15):1886-98. Genes Dev. 2000. PMID: 10921903 Free PMC article.
References
-
- Barlow, C., M. Liyanage, P.B. Moens, K. Nagahsima, K. Brown, S. Rottinghaus, S.P. Jackson, D. Tagle, T. Ried, and A. Wynshaw-Boris. 1998. ATM deficiency results in severe meiotic disruption as early as leptotenema in prophase I. Development (in press). - PubMed
-
- Baumann P, Benson FE, West SC. Human Rad51 promotes ATP-dependent homologous pairing and strand transfer reactions in vitro. Cell. 1996;87:757–766. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
