Identification and characterization of variants in PSEN1, PSEN2, and APP genes in Chinese patients with early-onset Alzheimer's disease
- PMID: 40016812
- PMCID: PMC11866898
- DOI: 10.1186/s13195-025-01702-0
Identification and characterization of variants in PSEN1, PSEN2, and APP genes in Chinese patients with early-onset Alzheimer's disease
Abstract
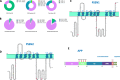
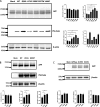
Variants in PSEN1, PSEN2, and APP are major genetic causes of early-onset Alzheimer's disease (EOAD). Our study aimed to identify the genotypic and phenotypic spectrums in a Chinese EOAD cohort and confirm their pathogenicity by functional analysis. This study included 304 unrelated clinically diagnosed EOAD participants of Chinese Han ancestry. Whole-exome sequencing revealed that 26 out of 304 individuals (8.6%) carried rare variants in PSEN1, PSEN2, and APP, including 16 in PSEN1 (5.3%), 6 in PSEN2 (2.0%), and 4 in APP (1.3%). Eight variants were novel, including PSEN1 p.Q56R, PSEN1 p.L174P, PSEN1 p.S289P, PSEN1 p.Y466C, PSEN2 p.R17W, PSEN2 p.F331Y, APP p.D197N, and APP p.D252V. Functional study revealed that the PS1 L174P, S289P, R377M, Y466C, PS2 V214L, and M239T mutants increased Aβ42 levels and Aβ42/Aβ40 ratios. The PS1 L174P, R377M, and Y466C mutants decreased the maturation of presenilin-1. Our findings highlight the prevalence and pathogenic significance of APP /PSENs variants in a Chinese EOAD cohort and expand the phenotypic and genotypic spectrum of EOAD.
Keywords: APP; PSEN1; PSEN2; Alzheimer’s disease; Amyloid β; EOAD.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: The study was approved by the Ethics Committees of the Xuanwu Hospital of Capital Medical University (approval number: 2020026), and it was carried out in compliance with the Declaration of Helsinki’s principles. Written informed consent was obtained from each patient or their guardian. Consent for publication: Written informed consent for publication was obtained from the guardian of each patient. Competing interests: The authors declare no competing interests.
Figures

References
-
- McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology. 1984;34:939–44. - PubMed
-
- Wu L, Rosa-Neto P, Hsiung GY, et al. Early-onset familial Alzheimer’s disease (EOFAD). Can J Neurol Sci. 2012;39:436–45. - PubMed
-
- Alzheimer’s A. 2009 Alzheimer’s disease facts and figures. Alzheimers Dement. 2009;5:234–70. - PubMed
-
- Cacace R, Sleegers K, Van Broeckhoven C. Molecular genetics of early-onset Alzheimer’s disease revisited. Alzheimers Dement. 2016;12:733–48. - PubMed
-
- Shi Z, Wang Y, Liu S, et al. Clinical and neuroimaging characterization of Chinese dementia patients with PSEN1 and PSEN2 mutations. Dement Geriatr Cogn Disord. 2015;39:32–40. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

