Controlled mechanochemical coupling of anti-junctions in DNA origami arrays
- PMID: 39256353
- PMCID: PMC11387415
- DOI: 10.1038/s41467-024-51721-y
Controlled mechanochemical coupling of anti-junctions in DNA origami arrays
Abstract
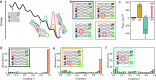
Allostery is a hallmark of cellular function and important in every biological system. Still, we are only starting to mimic it in the laboratory. Here, we introduce an approach to study aspects of allostery in artificial systems. We use a DNA origami domino array structure which-upon binding of trigger DNA strands-undergoes a stepwise allosteric conformational change. Using two FRET probes placed at specific positions in the DNA origami, we zoom in into single steps of this reaction cascade. Most of the steps are strongly coupled temporally and occur simultaneously. Introduction of activation energy barriers between different intermediate states alters this coupling and induces a time delay. We then apply these approaches to release a cargo DNA strand at a predefined step in the reaction cascade to demonstrate the applicability of this concept in tunable cascades of mechanochemical coupling with both spatial and temporal control.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Depressing time: Waiting, melancholia, and the psychoanalytic practice of care.In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. In: Kirtsoglou E, Simpson B, editors. The Time of Anthropology: Studies of Contemporary Chronopolitics. Abingdon: Routledge; 2020. Chapter 5. PMID: 36137063 Free Books & Documents. Review.
-
Sequence-Defined DNA Polymers: New Tools for DNA Nanotechnology and Nucleic Acid Therapy.Acc Chem Res. 2025 Jan 21;58(2):177-188. doi: 10.1021/acs.accounts.4c00580. Epub 2025 Jan 8. Acc Chem Res. 2025. PMID: 39772484
-
Dynamic Field Theory of Executive Function: Identifying Early Neurocognitive Markers.Monogr Soc Res Child Dev. 2024 Dec;89(3):7-109. doi: 10.1111/mono.12478. Monogr Soc Res Child Dev. 2024. PMID: 39628288 Free PMC article.
-
Double J Placement Methods Comparative Analysis.2023 May 30. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2023 May 30. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 29494060 Free Books & Documents.
-
Exploring conceptual and theoretical frameworks for nurse practitioner education: a scoping review protocol.JBI Database System Rev Implement Rep. 2015 Oct;13(10):146-55. doi: 10.11124/jbisrir-2015-2150. JBI Database System Rev Implement Rep. 2015. PMID: 26571290
References
MeSH terms
Substances
Grants and funding
- 03ZU1201AA/Bundesministerium für Bildung und Forschung (Federal Ministry of Education and Research)
- 1R35GM153472/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- RM1 GM145394/GM/NIGMS NIH HHS/United States
- ECCS-2328217/National Science Foundation (NSF)
- R35 GM153472/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources

