An acidophilic fungus promotes prey digestion in a carnivorous plant
- PMID: 39090391
- PMCID: PMC11445062
- DOI: 10.1038/s41564-024-01766-y
An acidophilic fungus promotes prey digestion in a carnivorous plant
Abstract
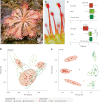
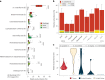
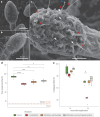
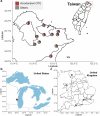
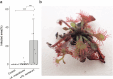
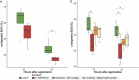
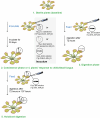
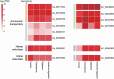
Leaves of the carnivorous sundew plants (Drosera spp.) secrete mucilage that hosts microorganisms, but whether this microbiota contributes to prey digestion is unclear. We identified the acidophilic fungus Acrodontium crateriforme as the dominant species in the mucilage microbial communities, thriving in multiple sundew species across the global range. The fungus grows and sporulates on sundew glands as its preferred acidic environment, and its presence in traps increased the prey digestion process. A. crateriforme has a reduced genome similar to other symbiotic fungi. During A. crateriforme-Drosera spatulata coexistence and digestion of prey insects, transcriptomes revealed significant gene co-option in both partners. Holobiont expression patterns during prey digestion further revealed synergistic effects in several gene families including fungal aspartic and sedolisin peptidases, facilitating prey digestion in leaves, as well as nutrient assimilation and jasmonate signalling pathway expression. This study establishes that botanical carnivory is defined by adaptations involving microbial partners and interspecies interactions.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures







Similar articles
-
Dynamic calcium signals mediate the feeding response of the carnivorous sundew plant.Proc Natl Acad Sci U S A. 2022 Jul 26;119(30):e2206433119. doi: 10.1073/pnas.2206433119. Epub 2022 Jul 12. Proc Natl Acad Sci U S A. 2022. PMID: 35858457 Free PMC article.
-
Wound and insect-induced jasmonate accumulation in carnivorous Drosera capensis: two sides of the same coin.Plant Biol (Stuttg). 2014 Sep;16(5):982-7. doi: 10.1111/plb.12148. Epub 2014 Feb 5. Plant Biol (Stuttg). 2014. PMID: 24499476
-
The role of electrical and jasmonate signalling in the recognition of captured prey in the carnivorous sundew plant Drosera capensis.New Phytol. 2017 Mar;213(4):1818-1835. doi: 10.1111/nph.14352. Epub 2016 Dec 9. New Phytol. 2017. PMID: 27933609
-
Jasmonate signalling in carnivorous plants: copycat of plant defence mechanisms.J Exp Bot. 2019 Jul 5;70(13):3379-3389. doi: 10.1093/jxb/erz188. J Exp Bot. 2019. PMID: 31120525 Review.
-
A novel insight into the cost-benefit model for the evolution of botanical carnivory.Ann Bot. 2015 Jun;115(7):1075-92. doi: 10.1093/aob/mcv050. Epub 2015 May 6. Ann Bot. 2015. PMID: 25948113 Free PMC article. Review.
References
-
- Darwin, C. & Darwin, F. Insectivorous Plants (J. Murray, 1888).
-
- Alessandri, G., Rizzo, S. M., Ossiprandi, M. C., van Sinderen, D. & Ventura, M. Creating an atlas to visualize the biodiversity of the mammalian gut microbiota. Curr. Opin. Biotechnol.73, 28–33 (2022). - PubMed
-
- Sirová, D. et al. Microbial community development in the traps of aquatic Utricularia species. Aquat. Bot.90, 129–136 (2009).
-
- Takeuchi, Y. et al. Bacterial diversity and composition in the fluid of pitcher plants of the genus Nepenthes. Syst. Appl. Microbiol.38, 330–339 (2015). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

