Autophagy and ALS: mechanistic insights and therapeutic implications
- PMID: 34057020
- PMCID: PMC8942428
- DOI: 10.1080/15548627.2021.1926656
Autophagy and ALS: mechanistic insights and therapeutic implications
Abstract
Mechanisms of protein homeostasis are crucial for overseeing the clearance of misfolded and toxic proteins over the lifetime of an organism, thereby ensuring the health of neurons and other cells of the central nervous system. The highly conserved pathway of autophagy is particularly necessary for preventing and counteracting pathogenic insults that may lead to neurodegeneration. In line with this, mutations in genes that encode essential autophagy factors result in impaired autophagy and lead to neurodegenerative conditions such as amyotrophic lateral sclerosis (ALS). However, the mechanistic details underlying the neuroprotective role of autophagy, neuronal resistance to autophagy induction, and the neuron-specific effects of autophagy-impairing mutations remain incompletely defined. Further, the manner and extent to which non-cell autonomous effects of autophagy dysfunction contribute to ALS pathogenesis are not fully understood. Here, we review the current understanding of the interplay between autophagy and ALS pathogenesis by providing an overview of critical steps in the autophagy pathway, with special focus on pivotal factors impaired by ALS-causing mutations, their physiologic effects on autophagy in disease models, and the cell type-specific mechanisms regulating autophagy in non-neuronal cells which, when impaired, can contribute to neurodegeneration. This review thereby provides a framework not only to guide further investigations of neuronal autophagy but also to refine therapeutic strategies for ALS and related neurodegenerative diseases.Abbreviations: ALS: amyotrophic lateral sclerosis; Atg: autophagy-related; CHMP2B: charged multivesicular body protein 2B; DPR: dipeptide repeat; FTD: frontotemporal dementia; iPSC: induced pluripotent stem cell; LIR: LC3-interacting region; MAP1LC3/LC3: microtubule associated protein 1 light chain 3; MTOR: mechanistic target of rapamycin kinase; PINK1: PTEN induced kinase 1; RNP: ribonuclear protein; sALS: sporadic ALS; SPHK1: sphingosine kinase 1; TARDBP/TDP-43: TAR DNA binding protein; TBK1: TANK-binding kinase 1; TFEB: transcription factor EB; ULK: unc-51 like autophagy activating kinase; UPR: unfolded protein response; UPS: ubiquitin-proteasome system; VCP: valosin containing protein.
Keywords: Amyotrophic lateral sclerosis; C9orf72; CHMP2B; SQSTM1/p62; TBK1; macroautophagy; mitophagy; myelinophagy; neuronal autophagy; optineurin.
Figures
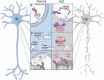
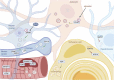
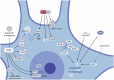
Similar articles
-
C9orf72 ALS-FTD: recent evidence for dysregulation of the autophagy-lysosome pathway at multiple levels.Autophagy. 2021 Nov;17(11):3306-3322. doi: 10.1080/15548627.2021.1872189. Epub 2021 Feb 26. Autophagy. 2021. PMID: 33632058 Free PMC article. Review.
-
PTK2/FAK regulates UPS impairment via SQSTM1/p62 phosphorylation in TARDBP/TDP-43 proteinopathies.Autophagy. 2020 Aug;16(8):1396-1412. doi: 10.1080/15548627.2019.1686729. Epub 2019 Nov 5. Autophagy. 2020. PMID: 31690171 Free PMC article.
-
Organelle-specific autophagy in inflammatory diseases: a potential therapeutic target underlying the quality control of multiple organelles.Autophagy. 2021 Feb;17(2):385-401. doi: 10.1080/15548627.2020.1725377. Epub 2020 Feb 12. Autophagy. 2021. PMID: 32048886 Free PMC article. Review.
-
Retinoic acid worsens ATG10-dependent autophagy impairment in TBK1-mutant hiPSC-derived motoneurons through SQSTM1/p62 accumulation.Autophagy. 2019 Oct;15(10):1719-1737. doi: 10.1080/15548627.2019.1589257. Epub 2019 Apr 2. Autophagy. 2019. PMID: 30939964 Free PMC article.
-
Lysosome dysfunction as a cause of neurodegenerative diseases: Lessons from frontotemporal dementia and amyotrophic lateral sclerosis.Neurobiol Dis. 2021 Jul;154:105360. doi: 10.1016/j.nbd.2021.105360. Epub 2021 Mar 31. Neurobiol Dis. 2021. PMID: 33812000 Free PMC article. Review.
Cited by
-
Autophagy: Identification of MTMR5 as a neuron-enriched suppressor.Curr Biol. 2022 Jun 20;32(12):R574-R577. doi: 10.1016/j.cub.2022.04.075. Curr Biol. 2022. PMID: 35728530 Free PMC article.
-
ALS-related p97 R155H mutation disrupts lysophagy in iPSC-derived motor neurons.Stem Cell Reports. 2024 Mar 12;19(3):366-382. doi: 10.1016/j.stemcr.2024.01.002. Epub 2024 Feb 8. Stem Cell Reports. 2024. PMID: 38335961 Free PMC article.
-
N-Acetylcysteine Inhibits Coxsackievirus B3 Replication by Downregulating Eukaryotic Translation Elongation Factor 1 Alpha 1.Viruses. 2024 Sep 23;16(9):1503. doi: 10.3390/v16091503. Viruses. 2024. PMID: 39339978 Free PMC article.
-
Extracellular Vesicles as Potential Biomarkers in Amyotrophic Lateral Sclerosis.Genes (Basel). 2023 Jan 27;14(2):325. doi: 10.3390/genes14020325. Genes (Basel). 2023. PMID: 36833252 Free PMC article. Review.
-
Potential Diets to Improve Mitochondrial Activity in Amyotrophic Lateral Sclerosis.Diseases. 2022 Dec 1;10(4):117. doi: 10.3390/diseases10040117. Diseases. 2022. PMID: 36547203 Free PMC article. Review.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
