Evaluation of extraction and amplification assays for the detection of SARS-CoV-2 at Auckland Hospital laboratory during the COVID-19 outbreak in New Zealand
- PMID: 33345831
- PMCID: PMC7837327
- DOI: 10.1016/j.jviromet.2020.114042
Evaluation of extraction and amplification assays for the detection of SARS-CoV-2 at Auckland Hospital laboratory during the COVID-19 outbreak in New Zealand
Abstract
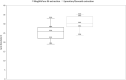
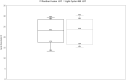
Utilising diverse molecular platforms has formed a solid foundation in New Zealand's COVID-19 response. We evaluated multiple extraction and PCR assays for the detection of SARS-CoV-2. We included 65 positive samples which were run on the Panther Fusion using a laboratory developed test (LDT, E gene target). Where viral RNA was extracted by MagNA Pure (MP) 96 extraction platform or EpMotion 5075/Geneaid extraction kit, SARS-CoV-2 detection was performed on Light Cycler (LC) 480 using a LDT (E gene) or 3 commercial assays; Certest Viasure (Orf1ab, N genes) GenePro (E, RdRp genes) and A* Star Fortitude (proprietary target). Median Cts on LC 480 LDT for specimens (n = 9) extracted on MP 96 (26.6) were lower than on EpMotion (31.6) whereas median Cts for specimens (n = 10) extracted on the Panther Fusion LDT (23.1) were comparable with MP 96 /LC480 LDT (23.6). Specimens tested on Panther Fusion LDT (n = 28), extracted by MP 96, and amplified using commercial assays showed good concordance with a few exceptions; lower median Ct values were seen for 2 targets on GenePro (16.9, 21.5) and Viasure (19.5, 21.1) than for the Panther Fusion LDT (24.2) and A* Star Fortitude (25.6). Specimens tested on MP 96 (n = 18) had comparable results using commercial assays, with lower median Cts for Viasure (22.2, 23.7) compared with the LC 480 LDT (24.7), GenePro (24.7,25.7) and A*Fortitude (25.1) assays. The study provides an early assessment of the performance characteristics of 3 extraction methods for viral RNA and 5 PCR assays for the detection of SARS-CoV-2.
Keywords: COVID-19; New Zealand; RT-PCR; SARS-CoV-2; Viral RNA extraction.
Copyright © 2020 Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors report no declarations of interest.
Figures
Similar articles
-
Multiplexing primer/probe sets for detection of SARS-CoV-2 by qRT-PCR.J Clin Virol. 2020 Aug;129:104499. doi: 10.1016/j.jcv.2020.104499. Epub 2020 Jun 8. J Clin Virol. 2020. PMID: 32535397 Free PMC article.
-
Use of the variplex™ SARS-CoV-2 RT-LAMP as a rapid molecular assay to complement RT-PCR for COVID-19 diagnosis.J Clin Virol. 2020 Nov;132:104616. doi: 10.1016/j.jcv.2020.104616. Epub 2020 Aug 31. J Clin Virol. 2020. PMID: 32891938 Free PMC article.
-
Development and implementation of a RT-qPCR extraction-free protocol for the detection of SARS-CoV-2 and impact on the turn-around-time.J Med Virol. 2021 Apr;93(4):2538-2542. doi: 10.1002/jmv.26782. Epub 2021 Jan 22. J Med Virol. 2021. PMID: 33415765
-
Summary of the Available Molecular Methods for Detection of SARS-CoV-2 during the Ongoing Pandemic.Int J Mol Sci. 2021 Jan 28;22(3):1298. doi: 10.3390/ijms22031298. Int J Mol Sci. 2021. PMID: 33525651 Free PMC article. Review.
-
Laboratory testing for the diagnosis of COVID-19.Biochem Biophys Res Commun. 2021 Jan 29;538:226-230. doi: 10.1016/j.bbrc.2020.10.069. Epub 2020 Oct 28. Biochem Biophys Res Commun. 2021. PMID: 33139015 Free PMC article. Review.
Cited by
-
Ultraviolet-C Irradiation, Heat, and Storage as Potential Methods of Inactivating SARS-CoV-2 and Bacterial Pathogens on Filtering Facepiece Respirators.Pathogens. 2022 Jan 10;11(1):83. doi: 10.3390/pathogens11010083. Pathogens. 2022. PMID: 35056031 Free PMC article.
-
COVID-19 deep classification network based on convolution and deconvolution local enhancement.Comput Biol Med. 2021 Aug;135:104588. doi: 10.1016/j.compbiomed.2021.104588. Epub 2021 Jun 22. Comput Biol Med. 2021. PMID: 34182330 Free PMC article.
-
Rapid Response to SARS-CoV-2 in Aotearoa New Zealand: Implementation of a Diagnostic Test and Characterization of the First COVID-19 Cases in the South Island.Viruses. 2021 Nov 4;13(11):2222. doi: 10.3390/v13112222. Viruses. 2021. PMID: 34835031 Free PMC article.
References
-
- Corman V.M., Landt O., Kaiser M., Molenkamp R., Meijer A., Chu D.K., Bleicker T., Brünink S., Schneider J., Schmidt M.L., Mulders D.G., Haagmans B.L., van der Veer B., van den Brink S., Wijsman L., Goderski G., Romette J.L., Ellis J., Zambon M., Peiris M., Drosten C. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020;25(3):2000045. doi: 10.2807/1560-7917.ES.2020.25.3.20000. - DOI - PMC - PubMed
-
- Johns Hopkins University . 2020. COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University.https://coronavirus.jhu.edu/map.html
-
- NZ Government . 2020. Ministry of Health Website for Case Definition.https://www.health.govt.nz/our-work/diseases-and-conditions/covid-19-nov...
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

