Cyclic di-AMP, a second messenger of primary importance: tertiary structures and binding mechanisms
- PMID: 32095817
- PMCID: PMC7102992
- DOI: 10.1093/nar/gkaa112
Cyclic di-AMP, a second messenger of primary importance: tertiary structures and binding mechanisms
Abstract
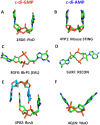
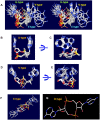
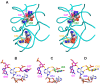
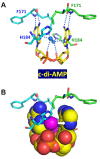
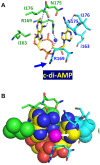
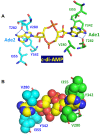
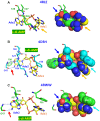
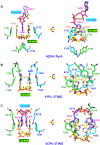
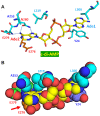
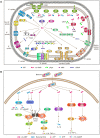
Cyclic diadenylate (c-di-AMP) is a widespread second messenger in bacteria and archaea that is involved in the maintenance of osmotic pressure, response to DNA damage, and control of central metabolism, biofilm formation, acid stress resistance, and other functions. The primary importance of c-di AMP stems from its essentiality for many bacteria under standard growth conditions and the ability of several eukaryotic proteins to sense its presence in the cell cytoplasm and trigger an immune response by the host cells. We review here the tertiary structures of the domains that regulate c-di-AMP synthesis and signaling, and the mechanisms of c-di-AMP binding, including the principal conformations of c-di-AMP, observed in various crystal structures. We discuss how these c-di-AMP molecules are bound to the protein and riboswitch receptors and what kinds of interactions account for the specific high-affinity binding of the c-di-AMP ligand. We describe seven kinds of non-covalent-π interactions between c-di-AMP and its receptor proteins, including π-π, C-H-π, cation-π, polar-π, hydrophobic-π, anion-π and the lone pair-π interactions. We also compare the mechanisms of c-di-AMP and c-di-GMP binding by the respective receptors that allow these two cyclic dinucleotides to control very different biological functions.
Published by Oxford University Press on behalf of Nucleic Acids Research 2020.
Figures
Similar articles
-
A jack of all trades: the multiple roles of the unique essential second messenger cyclic di-AMP.Mol Microbiol. 2015 Jul;97(2):189-204. doi: 10.1111/mmi.13026. Epub 2015 May 9. Mol Microbiol. 2015. PMID: 25869574 Review.
-
A decade of research on the second messenger c-di-AMP.FEMS Microbiol Rev. 2020 Nov 24;44(6):701-724. doi: 10.1093/femsre/fuaa019. FEMS Microbiol Rev. 2020. PMID: 32472931 Free PMC article. Review.
-
Nuclease-Resistant c-di-AMP Derivatives That Differentially Recognize RNA and Protein Receptors.Biochemistry. 2016 Feb 16;55(6):837-49. doi: 10.1021/acs.biochem.5b00965. Epub 2016 Feb 3. Biochemistry. 2016. PMID: 26789423 Free PMC article.
-
Structural Insights into the Distinct Binding Mode of Cyclic Di-AMP with SaCpaA_RCK.Biochemistry. 2015 Aug 11;54(31):4936-51. doi: 10.1021/acs.biochem.5b00633. Epub 2015 Jul 27. Biochemistry. 2015. PMID: 26171638
-
Making and Breaking of an Essential Poison: the Cyclases and Phosphodiesterases That Produce and Degrade the Essential Second Messenger Cyclic di-AMP in Bacteria.J Bacteriol. 2018 Dec 7;201(1):e00462-18. doi: 10.1128/JB.00462-18. Print 2019 Jan 1. J Bacteriol. 2018. PMID: 30224435 Free PMC article. Review.
Cited by
-
The many roles of cyclic di-AMP to control the physiology of Bacillus subtilis.Microlife. 2023 Oct 20;4:uqad043. doi: 10.1093/femsml/uqad043. eCollection 2023. Microlife. 2023. PMID: 37954098 Free PMC article. Review.
-
Gas and light: triggers of c-di-GMP-mediated regulation.FEMS Microbiol Rev. 2023 Jul 5;47(4):fuad034. doi: 10.1093/femsre/fuad034. FEMS Microbiol Rev. 2023. PMID: 37339911 Free PMC article. Review.
-
(p)ppGpp: Magic Modulators of Bacterial Physiology and Metabolism.Front Microbiol. 2020 Sep 7;11:2072. doi: 10.3389/fmicb.2020.02072. eCollection 2020. Front Microbiol. 2020. PMID: 33013756 Free PMC article.
-
Crystal structure and functional implication of a bacterial cyclic AMP-AMP-GMP synthetase.Nucleic Acids Res. 2021 May 7;49(8):4725-4737. doi: 10.1093/nar/gkab165. Nucleic Acids Res. 2021. PMID: 33836064 Free PMC article.
-
Structures of the DarR transcription regulator reveal unique modes of second messenger and DNA binding.Nat Commun. 2023 Nov 9;14(1):7239. doi: 10.1038/s41467-023-42823-0. Nat Commun. 2023. PMID: 37945601 Free PMC article.
References
-
- Hsu C.-Y.J., Dennis D., Oger J.. Synthesis and physical characterization of bis 3′→5′ cyclic dinucleotides (NpNp): RNA polymerase inhibitors. Nucleosides Nucleotides. 1985; 4:377–389.
-
- Ross P., Mayer R., Weinhouse H., Amikam D., Huggirat Y., Benziman M., de Vroom E., Fidder A., de Paus P., Sliedregt L.A. et al. .. The cyclic diguanylic acid regulatory system of cellulose synthesis in Acetobacter xylinum. Chemical synthesis and biological activity of cyclic nucleotide dimer, trimer, and phosphothioate derivatives. J. Biol. Chem. 1990; 265:18933–18943. - PubMed
-
- Witte G., Hartung S., Buttner K., Hopfner K.P.. Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates. Mol. Cell. 2008; 30:167–178. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

