Genetic and phenotypic dissection of 1q43q44 microdeletion syndrome and neurodevelopmental phenotypes associated with mutations in ZBTB18 and HNRNPU
- PMID: 28283832
- PMCID: PMC5360844
- DOI: 10.1007/s00439-017-1772-0
Genetic and phenotypic dissection of 1q43q44 microdeletion syndrome and neurodevelopmental phenotypes associated with mutations in ZBTB18 and HNRNPU
Abstract
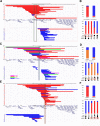
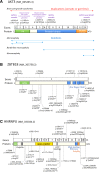
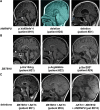
Subtelomeric 1q43q44 microdeletions cause a syndrome associating intellectual disability, microcephaly, seizures and anomalies of the corpus callosum. Despite several previous studies assessing genotype-phenotype correlations, the contribution of genes located in this region to the specific features of this syndrome remains uncertain. Among those, three genes, AKT3, HNRNPU and ZBTB18 are highly expressed in the brain and point mutations in these genes have been recently identified in children with neurodevelopmental phenotypes. In this study, we report the clinical and molecular data from 17 patients with 1q43q44 microdeletions, four with ZBTB18 mutations and seven with HNRNPU mutations, and review additional data from 37 previously published patients with 1q43q44 microdeletions. We compare clinical data of patients with 1q43q44 microdeletions with those of patients with point mutations in HNRNPU and ZBTB18 to assess the contribution of each gene as well as the possibility of epistasis between genes. Our study demonstrates that AKT3 haploinsufficiency is the main driver for microcephaly, whereas HNRNPU alteration mostly drives epilepsy and determines the degree of intellectual disability. ZBTB18 deletions or mutations are associated with variable corpus callosum anomalies with an incomplete penetrance. ZBTB18 may also contribute to microcephaly and HNRNPU to thin corpus callosum, but with a lower penetrance. Co-deletion of contiguous genes has additive effects. Our results confirm and refine the complex genotype-phenotype correlations existing in the 1qter microdeletion syndrome and define more precisely the neurodevelopmental phenotypes associated with genetic alterations of AKT3, ZBTB18 and HNRNPU in humans.
Conflict of interest statement
Funding
This study was financially supported by the Assistance Publique des Hôpitaux de Paris (AP-HP), PHRC (no PO81260), INSERM, Fondation Maladies Rares, Fondation de France (FdF—Engt no 15144), Agence de la Biomédecine, Agence Nationale de la Recherche (ANR Blanc CILAXCAL), and the “Investissements d’Avenir” programme ANR-10-IAIHU-06 (IHU-A-ICM). Dr Solveig Heide was supported by a master grant from the Fondation pour la Recherche Médicale (FRM). CD and CN are members of the Biopsy labex.
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
The study received approval from local ethical standards committees on human experimentation.
Informed consent
Informed written consent was obtained from each individual or their parents or legal representatives before blood sampling.
Figures
Similar articles
-
A de novo non-sense mutation in ZBTB18 in a patient with features of the 1q43q44 microdeletion syndrome.Eur J Hum Genet. 2014 Jun;22(6):844-6. doi: 10.1038/ejhg.2013.249. Epub 2013 Nov 6. Eur J Hum Genet. 2014. PMID: 24193349 Free PMC article.
-
A de novo nonsense mutation in ZBTB18 plus a de novo 15q13.3 microdeletion in a 6-year-old female.Am J Med Genet A. 2017 May;173(5):1251-1256. doi: 10.1002/ajmg.a.38145. Epub 2017 Mar 27. Am J Med Genet A. 2017. PMID: 28345786
-
High-resolution array CGH defines critical regions and candidate genes for microcephaly, abnormalities of the corpus callosum, and seizure phenotypes in patients with microdeletions of 1q43q44.Hum Genet. 2012 Jan;131(1):145-56. doi: 10.1007/s00439-011-1073-y. Epub 2011 Jul 29. Hum Genet. 2012. PMID: 21800092
-
Clinical findings of 21 previously unreported probands with HNRNPU-related syndrome and comprehensive literature review.Am J Med Genet A. 2020 Jul;182(7):1637-1654. doi: 10.1002/ajmg.a.61599. Epub 2020 Apr 22. Am J Med Genet A. 2020. PMID: 32319732 Review.
-
Expanding the phenotype of HNRNPU-related neurodevelopmental disorder with emphasis on seizure phenotype and review of literature.Am J Med Genet A. 2022 May;188(5):1497-1514. doi: 10.1002/ajmg.a.62677. Epub 2022 Feb 9. Am J Med Genet A. 2022. PMID: 35138025 Free PMC article. Review.
Cited by
-
Deficiency of the Heterogeneous Nuclear Ribonucleoprotein U locus leads to delayed hindbrain neurogenesis.Biol Open. 2023 Oct 15;12(10):bio060113. doi: 10.1242/bio.060113. Epub 2023 Oct 10. Biol Open. 2023. PMID: 37815090 Free PMC article.
-
HNRNPU: Key to Neurodevelopmental Disorders such as Intellectual Delay, Epilepsy, and Autism.Mol Syndromol. 2019 Jan;9(6):275-278. doi: 10.1159/000495204. Epub 2018 Dec 1. Mol Syndromol. 2019. PMID: 30800042 Free PMC article. No abstract available.
-
Understanding the impact of ZBTB18 missense variation on transcription factor function in neurodevelopment and disease.J Neurochem. 2022 May;161(3):219-235. doi: 10.1111/jnc.15572. Epub 2022 Feb 18. J Neurochem. 2022. PMID: 35083747 Free PMC article. Review.
-
Identification of an atypical microdeletion generating the RNF135-SUZ12 chimeric gene and causing a position effect in an NF1 patient with overgrowth.Hum Genet. 2017 Oct;136(10):1329-1339. doi: 10.1007/s00439-017-1832-5. Epub 2017 Aug 3. Hum Genet. 2017. PMID: 28776093
-
hnRNPs: roles in neurodevelopment and implication for brain disorders.Front Mol Neurosci. 2024 Jul 17;17:1411639. doi: 10.3389/fnmol.2024.1411639. eCollection 2024. Front Mol Neurosci. 2024. PMID: 39086926 Free PMC article. Review.
References
-
- Ballif BC, Rosenfeld JA, Traylor R, Theisen A, Bader PI, Ladda RL, Sell SL, Steinraths M, Surti U, McGuire M, Williams S, Farrell SA, Filiano J, Schnur RE, Coffey LB, Tervo RC, Stroud T, Marble M, Netzloff M, Hanson K, Aylsworth AS, Bamforth JS, Babu D, Niyazov DM, Ravnan JB, Schultz RA, Lamb AN, Torchia BS, Bejjani BA, Shaffer LG. High-resolution array CGH defines critical regions and candidate genes for microcephaly, abnormalities of the corpus callosum, and seizure phenotypes in patients with microdeletions of 1q43q44. Hum Genet. 2012;131:145–156. doi: 10.1007/s00439-011-1073-y. - DOI - PubMed
-
- Boland E, Clayton-Smith J, Woo VG, McKee S, Manson FD, Medne L, Zackai E, Swanson EA, Fitzpatrick D, Millen KJ, Sherr EH, Dobyns WB, Black GC. Mapping of deletion and translocation breakpoints in 1q44 implicates the serine/threonine kinase AKT3 in postnatal microcephaly and agenesis of the corpus callosum. Am J Hum Genet. 2007;81:292–303. doi: 10.1086/519999. - DOI - PMC - PubMed
-
- Caliebe A, Kroes HY, van der Smagt JJ, Martin-Subero JI, Tonnies H, van’t Slot R, Nievelstein RA, Muhle H, Stephani U, Alfke K, Stefanova I, Hellenbroich Y, Gillessen-Kaesbach G, Hochstenbach R, Siebert R, Poot M. Four patients with speech delay, seizures and variable corpus callosum thickness sharing a 0.440 Mb deletion in region 1q44 containing the HNRPU gene. Eur J Med Genet. 2010;53:179–185. doi: 10.1016/j.ejmg.2010.04.001. - DOI - PubMed
-
- Carvill GL, Heavin SB, Yendle SC, McMahon JM, O’Roak BJ, Cook J, Khan A, Dorschner MO, Weaver M, Calvert S, Malone S, Wallace G, Stanley T, Bye AM, Bleasel A, Howell KB, Kivity S, Mackay MT, Rodriguez-Casero V, Webster R, Korczyn A, Afawi Z, Zelnick N, Lerman-Sagie T, Lev D, Moller RS, Gill D, Andrade DM, Freeman JL, Sadleir LG, Shendure J, Berkovic SF, Scheffer IE, Mefford HC. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat Genet. 2013;45:825–830. doi: 10.1038/ng.2646. - DOI - PMC - PubMed
Urls/Resources
-
- BIOBASE HGMD Professional: http://www.biobase-international.com/product/hgmd
-
- Exome Variant Server: http://evs.gs.washington.edu/EVS/
-
- ExAC Browser (Beta) | Exome Aggregation Consortium: http://exac.broadinstitute.org/
-
- GeneMatcher: https://genematcher.org/
-
- NCBI Pubmed: http://www.ncbi.nlm.nih.gov/pubmed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

