Biogeography of Oenococcus oeni Reveals Distinctive but Nonspecific Populations in Wine-Producing Regions
- PMID: 27864168
- PMCID: PMC5244301
- DOI: 10.1128/AEM.02322-16
Biogeography of Oenococcus oeni Reveals Distinctive but Nonspecific Populations in Wine-Producing Regions
Abstract
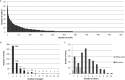
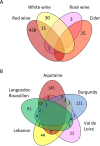
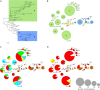
Understanding the mechanisms behind the typicity of regional wines inevitably brings attention to microorganisms associated with their production. Oenococcus oeni is the main bacterial species involved in wine and cider making. It develops after the yeast-driven alcoholic fermentation and performs the malolactic fermentation, which improves the taste and aromatic complexity of most wines. Here, we have evaluated the diversity and specificity of O. oeni strains in six regions. A total of 235 wines and ciders were collected during spontaneous malolactic fermentations and used to isolate 3,212 bacterial colonies. They were typed by multilocus variable analysis, which disclosed a total of 514 O. oeni strains. Their phylogenetic relationships were evaluated by a second typing method based on single nucleotide polymorphism (SNP) analysis. Taken together, the results indicate that each region holds a high diversity of strains that constitute a unique population. However, strains present in each region belong to diverse phylogenetic groups, and the same groups can be detected in different regions, indicating that strains are not genetically adapted to regions. In contrast, greater strain identity was seen for cider, white wine, or red wine of Burgundy, suggesting that genetic adaptation to these products occurred.
Importance: This study reports the isolation, genotyping, and geographic distribution analysis of the largest collection of O. oeni strains performed to date. It reveals that there is very high diversity of strains in each region, the majority of them being detected in a single region. The study also reports the development of an SNP genotyping method that is useful for analyzing the distribution of O. oeni phylogroups. The results show that strains are not genetically adapted to regions but to specific types of wines. They reveal new phylogroups of strains, particularly two phylogroups associated with white wines and red wines of Burgundy. Taken together, the results shed light on the diversity and specificity of wild strains of O. oeni, which is crucial for understanding their real contribution to the unique properties of wines.
Keywords: Oenococcus oeni; biogeography; lactic acid bacteria; wine.
Copyright © 2017 American Society for Microbiology.
Figures
Similar articles
-
Phylogenomic Analysis of Oenococcus oeni Reveals Specific Domestication of Strains to Cider and Wines.Genome Biol Evol. 2015 May 14;7(6):1506-18. doi: 10.1093/gbe/evv084. Genome Biol Evol. 2015. PMID: 25977455 Free PMC article.
-
Alcoholic fermentation drives the selection of Oenococcus oeni strains in wine but not in cider.Int J Food Microbiol. 2023 Sep 2;400:110276. doi: 10.1016/j.ijfoodmicro.2023.110276. Epub 2023 May 29. Int J Food Microbiol. 2023. PMID: 37270987
-
Characterization and technological properties of Oenococcus oeni strains from wine spontaneous malolactic fermentations: a framework for selection of new starter cultures.J Appl Microbiol. 2010 Jan;108(1):285-98. doi: 10.1111/j.1365-2672.2009.04428.x. J Appl Microbiol. 2010. PMID: 19614854
-
Distribution of Oenococcus oeni populations in natural habitats.Appl Microbiol Biotechnol. 2019 Apr;103(7):2937-2945. doi: 10.1007/s00253-019-09689-z. Epub 2019 Feb 20. Appl Microbiol Biotechnol. 2019. PMID: 30788540 Free PMC article. Review.
-
Oenococcus oeni and the genomic era.FEMS Microbiol Rev. 2017 Aug 1;41(Supp_1):S84-S94. doi: 10.1093/femsre/fux034. FEMS Microbiol Rev. 2017. PMID: 28830095 Review.
Cited by
-
From the Vineyard to the Winery: How Microbial Ecology Drives Regional Distinctiveness of Wine.Front Microbiol. 2019 Nov 20;10:2679. doi: 10.3389/fmicb.2019.02679. eCollection 2019. Front Microbiol. 2019. PMID: 31824462 Free PMC article. Review.
-
Exopolysaccharides Producing Lactic Acid Bacteria in Wine and Other Fermented Beverages: For Better or for Worse?Foods. 2021 Sep 17;10(9):2204. doi: 10.3390/foods10092204. Foods. 2021. PMID: 34574312 Free PMC article. Review.
-
Microorganisms in Fermented Apple Beverages: Current Knowledge and Future Directions.Microorganisms. 2017 Jul 25;5(3):39. doi: 10.3390/microorganisms5030039. Microorganisms. 2017. PMID: 28757560 Free PMC article. Review.
-
Distribution of Prophages in the Oenococcus oeni Species.Microorganisms. 2021 Apr 16;9(4):856. doi: 10.3390/microorganisms9040856. Microorganisms. 2021. PMID: 33923461 Free PMC article.
-
Mapping the Physiological Response of Oenococcus oeni to Ethanol Stress Using an Extended Genome-Scale Metabolic Model.Front Microbiol. 2018 Mar 1;9:291. doi: 10.3389/fmicb.2018.00291. eCollection 2018. Front Microbiol. 2018. PMID: 29545779 Free PMC article.
References
-
- Martiny JBH, Bohannan BJM, Brown JH, Colwell RK, Fuhrman JA, Green JL, Horner-Devine MC, Kane M, Krumins JA, Kuske CR, Morin PJ, Naeem S, Øvreås L, Reysenbach A-L, Smith VH, Staley JT. 2006. Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4:102–112. doi:10.1038/nrmicro1341. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

