Two-Component Signal Transduction Systems That Regulate the Temporal and Spatial Expression of Myxococcus xanthus Sporulation Genes
- PMID: 26369581
- PMCID: PMC4719452
- DOI: 10.1128/JB.00474-15
Two-Component Signal Transduction Systems That Regulate the Temporal and Spatial Expression of Myxococcus xanthus Sporulation Genes
Abstract
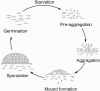
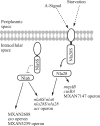
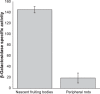
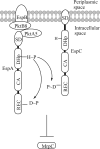
When starved for nutrients, Myxococcus xanthus produces a biofilm that contains a mat of rod-shaped cells, known as peripheral rods, and aerial structures called fruiting bodies, which house thousands of dormant and stress-resistant spherical spores. Because rod-shaped cells differentiate into spherical, stress-resistant spores and spore differentiation occurs only in nascent fruiting bodies, many genes and multiple levels of regulation are required. Over the past 2 decades, many regulators of the temporal and spatial expression of M. xanthus sporulation genes have been uncovered. Of these sporulation gene regulators, two-component signal transduction circuits, which typically contain a histidine kinase sensor protein and a transcriptional regulator known as response regulator, are among the best characterized. In this review, we discuss prototypical two-component systems (Nla6S/Nla6 and Nla28S/Nla28) that regulate an early, preaggregation phase of sporulation gene expression during fruiting body development. We also discuss orphan response regulators (ActB and FruA) that regulate a later phase of sporulation gene expression, which begins during the aggregation stage of fruiting body development. In addition, we summarize the research on a complex two-component system (Esp) that is important for the spatial regulation of sporulation.
Copyright © 2016, American Society for Microbiology. All Rights Reserved.
Figures
Similar articles
-
The enhancer binding protein Nla6 regulates developmental genes that are important for Myxococcus xanthus sporulation.J Bacteriol. 2015 Apr;197(7):1276-87. doi: 10.1128/JB.02408-14. Epub 2015 Feb 2. J Bacteriol. 2015. PMID: 25645554 Free PMC article.
-
The Nla28S/Nla28 two-component signal transduction system regulates sporulation in Myxococcus xanthus.J Bacteriol. 2012 Sep;194(17):4698-708. doi: 10.1128/JB.00225-12. Epub 2012 Jun 29. J Bacteriol. 2012. PMID: 22753068 Free PMC article.
-
TodK, a putative histidine protein kinase, regulates timing of fruiting body morphogenesis in Myxococcus xanthus.J Bacteriol. 2003 Sep;185(18):5452-64. doi: 10.1128/JB.185.18.5452-5464.2003. J Bacteriol. 2003. PMID: 12949097 Free PMC article.
-
Dual regulation with Ser/Thr kinase cascade and a His/Asp TCS in Myxococcus xanthus.Adv Exp Med Biol. 2008;631:111-21. doi: 10.1007/978-0-387-78885-2_7. Adv Exp Med Biol. 2008. PMID: 18792684 Review.
-
Molecular Mechanisms of Signaling in Myxococcus xanthus Development.J Mol Biol. 2016 Sep 25;428(19):3805-30. doi: 10.1016/j.jmb.2016.07.008. Epub 2016 Jul 16. J Mol Biol. 2016. PMID: 27430596 Review.
Cited by
-
Suppressor mutations reveal an NtrC-like response regulator, NmpR, for modulation of Type-IV Pili-dependent motility in Myxococcus xanthus.PLoS Genet. 2018 Oct 22;14(10):e1007714. doi: 10.1371/journal.pgen.1007714. eCollection 2018 Oct. PLoS Genet. 2018. PMID: 30346960 Free PMC article.
-
MifS, a DctB family histidine kinase, is a specific regulator of α-ketoglutarate response in Pseudomonas aeruginosa PAO1.Microbiology (Reading). 2020 Sep;166(9):867-879. doi: 10.1099/mic.0.000943. Microbiology (Reading). 2020. PMID: 32553056 Free PMC article.
-
The σ54 system directly regulates bacterial natural product genes.Sci Rep. 2021 Feb 26;11(1):4771. doi: 10.1038/s41598-021-84057-4. Sci Rep. 2021. PMID: 33637792 Free PMC article.
-
Transcriptional factor OmpR positively regulates prodigiosin biosynthesis in Serratia marcescens FZSF02 by binding with the promoter of the prodigiosin cluster.Front Microbiol. 2022 Nov 17;13:1041146. doi: 10.3389/fmicb.2022.1041146. eCollection 2022. Front Microbiol. 2022. PMID: 36466667 Free PMC article.
-
A Genomic Survey of Signalling in the Myxococcaceae.Microorganisms. 2020 Nov 6;8(11):1739. doi: 10.3390/microorganisms8111739. Microorganisms. 2020. PMID: 33171896 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

