devI is an evolutionarily young negative regulator of Myxococcus xanthus development
- PMID: 25645563
- PMCID: PMC4352663
- DOI: 10.1128/JB.02542-14
devI is an evolutionarily young negative regulator of Myxococcus xanthus development
Abstract
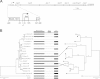
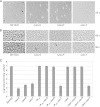
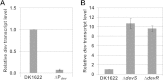
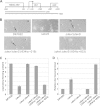
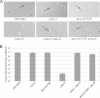
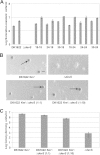
During starvation-induced development of Myxococcus xanthus, thousands of rod-shaped cells form mounds in which they differentiate into spores. The dev locus includes eight genes followed by clustered regularly interspaced short palindromic repeats (CRISPRs), comprising a CRISPR-Cas system (Cas stands for CRISPR associated) typically involved in RNA interference. Mutations in devS or devR of a lab reference strain permit mound formation but impair sporulation. We report that natural isolates of M. xanthus capable of normal development are highly polymorphic in the promoter region of the dev operon. We show that the dev promoter is predicted to be nonfunctional in most natural isolates and is dispensable for development of a laboratory reference strain. Moreover, deletion of the dev promoter or the small gene immediately downstream of it, here designated devI (development inhibitor), suppressed the sporulation defect of devS or devR mutants in the lab strain. Complementation experiments and the result of introducing a premature stop codon in devI support a model in which DevRS proteins negatively autoregulate expression of devI, whose 40-residue protein product DevI inhibits sporulation if overexpressed. DevI appears to act in a cell-autonomous manner since experiments with conditioned medium and with cell mixtures gave no indication of extracellular effects. Strikingly, we report that devI is entirely absent from most M. xanthus natural isolates and was only recently integrated into the developmental programs of some lineages. These results provide important new insights into both the evolutionary history of the dev operon and its mechanistic role in M. xanthus sporulation.
Importance: Certain mutations in the dev CRISPR-Cas (clustered regularly interspaced short palindromic repeat-associated) system of Myxococcus xanthus impair sporulation. The link between development and a CRISPR-Cas system has been a mystery. Surprisingly, DNA sequencing of natural isolates revealed that many appear to lack a functional dev promoter, yet these strains sporulate normally. Deletion of the dev promoter or the small gene downstream of it suppressed the sporulation defect of a lab strain with mutations in dev genes encoding Cas proteins. The results support a model in which the Cas proteins DevRS prevent overexpression of the small gene devI, which codes for an inhibitor of sporulation. Phylogenetic analysis of natural isolates suggests that devI and the dev promoter were only recently acquired in some lineages.
Copyright © 2015, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
The dev Operon Regulates the Timing of Sporulation during Myxococcus xanthus Development.J Bacteriol. 2017 Apr 25;199(10):e00788-16. doi: 10.1128/JB.00788-16. Print 2017 May 15. J Bacteriol. 2017. PMID: 28264995 Free PMC article.
-
Regulation of dev, an operon that includes genes essential for Myxococcus xanthus development and CRISPR-associated genes and repeats.J Bacteriol. 2007 May;189(10):3738-50. doi: 10.1128/JB.00187-07. Epub 2007 Mar 16. J Bacteriol. 2007. PMID: 17369305 Free PMC article.
-
devRS, an autoregulated and essential genetic locus for fruiting body development in Myxococcus xanthus.J Bacteriol. 1993 Nov;175(22):7450-62. doi: 10.1128/jb.175.22.7450-7462.1993. J Bacteriol. 1993. PMID: 7693658 Free PMC article.
-
Two-Component Signal Transduction Systems That Regulate the Temporal and Spatial Expression of Myxococcus xanthus Sporulation Genes.J Bacteriol. 2015 Sep 14;198(3):377-85. doi: 10.1128/JB.00474-15. Print 2016 Feb 1. J Bacteriol. 2015. PMID: 26369581 Free PMC article. Review.
-
Dual regulation with Ser/Thr kinase cascade and a His/Asp TCS in Myxococcus xanthus.Adv Exp Med Biol. 2008;631:111-21. doi: 10.1007/978-0-387-78885-2_7. Adv Exp Med Biol. 2008. PMID: 18792684 Review.
Cited by
-
Myxobacterial Genomics and Post-Genomics: A Review of Genome Biology, Genome Sequences and Related 'Omics Studies.Microorganisms. 2021 Oct 13;9(10):2143. doi: 10.3390/microorganisms9102143. Microorganisms. 2021. PMID: 34683464 Free PMC article. Review.
-
Highly Signal-Responsive Gene Regulatory Network Governing Myxococcus Development.Trends Genet. 2017 Jan;33(1):3-15. doi: 10.1016/j.tig.2016.10.006. Epub 2016 Dec 2. Trends Genet. 2017. PMID: 27916428 Free PMC article. Review.
-
Allopatric divergence of cooperators confers cheating resistance and limits effects of a defector mutation.BMC Ecol Evol. 2022 Dec 12;22(1):141. doi: 10.1186/s12862-022-02094-7. BMC Ecol Evol. 2022. PMID: 36510120 Free PMC article.
-
Killer prey: Ecology reverses bacterial predation.PLoS Biol. 2024 Jan 23;22(1):e3002454. doi: 10.1371/journal.pbio.3002454. eCollection 2024 Jan. PLoS Biol. 2024. PMID: 38261596 Free PMC article.
-
Alternative functions of CRISPR-Cas systems in the evolutionary arms race.Nat Rev Microbiol. 2022 Jun;20(6):351-364. doi: 10.1038/s41579-021-00663-z. Epub 2022 Jan 6. Nat Rev Microbiol. 2022. PMID: 34992260 Review.
References
-
- Velicer GJ, Mendes-Soares H, Wielgloss S. 2014. Whence comes social diversity? Ecological and evolutionary analysis of the myxobacteria, p 1–29. In Yang Z, Higgs P (ed), Myxobacteria: genomics, cellular and molecular biology. Caister Academic Press, Norfolk, United Kingdom.
-
- Munoz-Dorado J, Higgs P, Elias-Arnanz M. 2014. Abundance and complexity of signaling mechanisms in myxobacteria, p 127–149. In Yang Z, Higgs P (ed), Myxobacteria: genomics, cellular and molecular biology. Caister Academic Press, Norfolk, United Kingdom.
-
- Rajagopalan R, Sarwar Z, Garza AG, Kroos L. 2014. Developmental gene regulation, p 105–126. In Yang Z, Higgs P (ed), Myxobacteria: genomics, cellular and molecular biology. Caister Academic Press, Norfolk, United Kingdom.
-
- Higgs P, Hartzell PL, Holkenbrink C, Hoiczyk E. 2014. Myxococcus xanthus vegetative and developmental cell heterogeneity, p 51–77. In Yang Z, Higgs P (ed), Myxobacteria: genomics, cellular and molecular biology. Caister Academic Press, Norfolk, United Kingdom.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

