Defects in mitochondrial fatty acid synthesis result in failure of multiple aspects of mitochondrial biogenesis in Saccharomyces cerevisiae
- PMID: 24102902
- PMCID: PMC4153884
- DOI: 10.1111/mmi.12402
Defects in mitochondrial fatty acid synthesis result in failure of multiple aspects of mitochondrial biogenesis in Saccharomyces cerevisiae
Abstract
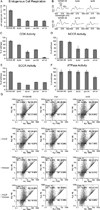
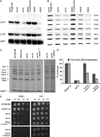
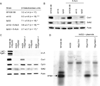
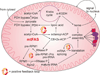
Mitochondrial fatty acid synthesis (mtFAS) shares acetyl-CoA with the Krebs cycle as a common substrate and is required for the production of octanoic acid (C8) precursors of lipoic acid (LA) in mitochondria. MtFAS is a conserved pathway essential for respiration. In a genetic screen in Saccharomyces cerevisiae designed to further elucidate the physiological role of mtFAS, we isolated mutants with defects in mitochondrial post-translational gene expression processes, indicating a novel link to mitochondrial gene expression and respiratory chain biogenesis. In our ensuing analysis, we show that mtFAS, but not lipoylation per se, is required for respiratory competence. We demonstrate that mtFAS is required for mRNA splicing, mitochondrial translation and respiratory complex assembly, and provide evidence that not LA per se, but fatty acids longer than C8 play a role in these processes. We also show that mtFAS- and LA-deficient strains suffer from a mild haem deficiency that may contribute to the respiratory complex assembly defect. Based on our data and previously published information, we propose a model implicating mtFAS as a sensor for mitochondrial acetyl-CoA availability and a co-ordinator of nuclear and mitochondrial gene expression by adapting the mitochondrial compartment to changes in the metabolic status of the cell.
© 2013 John Wiley & Sons Ltd.
Figures



Similar articles
-
ACP Acylation Is an Acetyl-CoA-Dependent Modification Required for Electron Transport Chain Assembly.Mol Cell. 2018 Aug 16;71(4):567-580.e4. doi: 10.1016/j.molcel.2018.06.039. Mol Cell. 2018. PMID: 30118679 Free PMC article.
-
Genetic dissection of the mitochondrial lipoylation pathway in yeast.BMC Biol. 2021 Jan 25;19(1):14. doi: 10.1186/s12915-021-00951-3. BMC Biol. 2021. PMID: 33487163 Free PMC article.
-
An engineered variant of MECR reductase reveals indispensability of long-chain acyl-ACPs for mitochondrial respiration.Nat Commun. 2023 Feb 4;14(1):619. doi: 10.1038/s41467-023-36358-7. Nat Commun. 2023. PMID: 36739436 Free PMC article.
-
Mitochondrial Fatty Acids and Neurodegenerative Disorders.Neuroscientist. 2021 Apr;27(2):143-158. doi: 10.1177/1073858420936162. Epub 2020 Jul 9. Neuroscientist. 2021. PMID: 32644907 Review.
-
Impact of Mitochondrial Fatty Acid Synthesis on Mitochondrial Biogenesis.Curr Biol. 2018 Oct 22;28(20):R1212-R1219. doi: 10.1016/j.cub.2018.08.022. Curr Biol. 2018. PMID: 30352195 Free PMC article. Review.
Cited by
-
A defect in mitochondrial fatty acid synthesis impairs iron metabolism and causes elevated ceramide levels.Nat Metab. 2023 Sep;5(9):1595-1614. doi: 10.1038/s42255-023-00873-0. Epub 2023 Aug 31. Nat Metab. 2023. PMID: 37653044 Free PMC article.
-
Lipoic acid biosynthesis defects.J Inherit Metab Dis. 2014 Jul;37(4):553-63. doi: 10.1007/s10545-014-9705-8. Epub 2014 Apr 29. J Inherit Metab Dis. 2014. PMID: 24777537 Review.
-
Expression and analysis of the SAM-dependent RNA methyltransferase Rsm22 from Saccharomyces cerevisiae.Acta Crystallogr D Struct Biol. 2021 Jun 1;77(Pt 6):840-853. doi: 10.1107/S2059798321004149. Epub 2021 May 19. Acta Crystallogr D Struct Biol. 2021. PMID: 34076597 Free PMC article.
-
Regulating the metabolic flux of pyruvate dehydrogenase bypass to enhance lipid production in Saccharomyces cerevisiae.Commun Biol. 2024 Oct 26;7(1):1399. doi: 10.1038/s42003-024-07103-7. Commun Biol. 2024. PMID: 39462103 Free PMC article.
-
Alternative reactions at the interface of glycolysis and citric acid cycle in Saccharomyces cerevisiae.FEMS Yeast Res. 2016 May;16(3):fow017. doi: 10.1093/femsyr/fow017. Epub 2016 Feb 18. FEMS Yeast Res. 2016. PMID: 26895788 Free PMC article.
References
-
- Adams A, Gottschling DE, Kaiser CA, Stearns T. Methods in yeast genetics. Cold Spring Harbor: Cold Spring Harbor Laboratory Press; 1997.
-
- Amiott EA, Jaehning JA. Sensitivity of the yeast mitochondrial RNA polymerase to +1 and +2 initiating nucleotides. J Biol Chem. 2006a;281:34982–34988. - PubMed
-
- Amiott EA, Jaehning JA. Mitochondrial transcription is regulated via an ATP "sensing" mechanism that couples RNA abundance to respiration. Mol Cell. 2006b;22:329–338. - PubMed
-
- Ausubel FM, Brent RE, Kingston DD, Moore JG, Seidman JA, editors. Current protocols in molecular biology. New York, NY: John Wiley and Sons, Inc; 1989. Current protocols in molecular biology.
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

