Flexible long-range loops in the VH gene region of the Igh locus facilitate the generation of a diverse antibody repertoire
- PMID: 23973221
- PMCID: PMC4810778
- DOI: 10.1016/j.immuni.2013.08.011
Flexible long-range loops in the VH gene region of the Igh locus facilitate the generation of a diverse antibody repertoire
Abstract
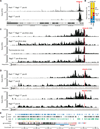
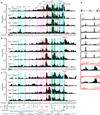
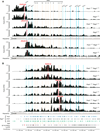
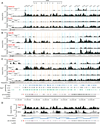
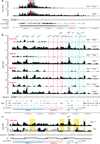
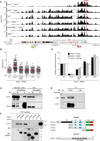
The immunoglobulin heavy-chain (Igh) locus undergoes large-scale contraction in pro-B cells, which facilitates VH-DJH recombination by juxtaposing distal VH genes next to the DJH-rearranged gene segment in the 3' proximal Igh domain. By using high-resolution mapping of long-range interactions, we demonstrate that local interaction domains established the three-dimensional structure of the extended Igh locus in lymphoid progenitors. In pro-B cells, these local domains engaged in long-range interactions across the Igh locus, which depend on the regulators Pax5, YY1, and CTCF. The large VH gene cluster underwent flexible long-range interactions with the more rigidly structured proximal domain, which probably ensures similar participation of all VH genes in VH-DJH recombination to generate a diverse antibody repertoire. These long-range interactions appear to be an intrinsic feature of the VH gene cluster, because they are still generated upon mutation of the Eμ enhancer, IGCR1 insulator, or 3' regulatory region in the proximal Igh domain.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures

Comment in
-
4C-ing the Igh Landscape.Immunity. 2013 Aug 22;39(2):199-201. doi: 10.1016/j.immuni.2013.08.014. Immunity. 2013. PMID: 23973215 Free PMC article.
Similar articles
-
Epigenetic modifications of the VH region after DJH recombination in Pro-B cells.Immunology. 2017 Oct;152(2):218-231. doi: 10.1111/imm.12758. Epub 2017 Jun 23. Immunology. 2017. PMID: 28502113 Free PMC article.
-
Sequential Enhancer Sequestration Dysregulates Recombination Center Formation at the IgH Locus.Mol Cell. 2018 Apr 5;70(1):21-33.e6. doi: 10.1016/j.molcel.2018.02.020. Epub 2018 Mar 22. Mol Cell. 2018. PMID: 29576529 Free PMC article.
-
IL-7R signaling activates widespread VH and DH gene usage to drive antibody diversity in bone marrow B cells.Cell Rep. 2021 Jul 13;36(2):109349. doi: 10.1016/j.celrep.2021.109349. Cell Rep. 2021. PMID: 34260907 Free PMC article.
-
Spatial Regulation of V-(D)J Recombination at Antigen Receptor Loci.Adv Immunol. 2015;128:93-121. doi: 10.1016/bs.ai.2015.07.006. Epub 2015 Aug 13. Adv Immunol. 2015. PMID: 26477366 Review.
-
Pax5: a master regulator of B cell development and leukemogenesis.Adv Immunol. 2011;111:179-206. doi: 10.1016/B978-0-12-385991-4.00005-2. Adv Immunol. 2011. PMID: 21970955 Review.
Cited by
-
CTCF and Its Partners: Shaper of 3D Genome during Development.Genes (Basel). 2022 Aug 2;13(8):1383. doi: 10.3390/genes13081383. Genes (Basel). 2022. PMID: 36011294 Free PMC article. Review.
-
YY1 control of mitochondrial-related genes does not account for regulation of immunoglobulin class switch recombination in mice.Eur J Immunol. 2020 Jun;50(6):822-838. doi: 10.1002/eji.201948385. Epub 2020 Mar 16. Eur J Immunol. 2020. PMID: 32092784 Free PMC article.
-
CTCF-Binding Elements Mediate Accessibility of RAG Substrates During Chromatin Scanning.Cell. 2018 Jun 28;174(1):102-116.e14. doi: 10.1016/j.cell.2018.04.035. Epub 2018 May 24. Cell. 2018. PMID: 29804837 Free PMC article.
-
Epigenetic Regulation of Individual Modules of the immunoglobulin heavy chain locus 3' Regulatory Region.Front Immunol. 2014 Apr 21;5:163. doi: 10.3389/fimmu.2014.00163. eCollection 2014. Front Immunol. 2014. PMID: 24795714 Free PMC article. Review.
-
3D trajectories adopted by coding and regulatory DNA elements: first-passage times for genomic interactions.Cell. 2014 Jul 17;158(2):339-352. doi: 10.1016/j.cell.2014.05.036. Epub 2014 Jul 3. Cell. 2014. PMID: 24998931 Free PMC article.
References
-
- Afshar R, Pierce S, Bolland DJ, Corcoran A, Oltz EM. Regulation of IgH gene assembly: role of the intronic enhancer and 5’DQ52 region in targeting DHJH recombination. J. Immunol. 2006;176:2439–2447. - PubMed
-
- Bolland DJ, Wood AL, Johnston CM, Bunting SF, Morgan G, Chakalova L, Fraser PJ, Corcoran AE. Antisense intergenic transcription in V(D)J recombination. Nat. Immunol. 2004;5:630–637. - PubMed
-
- Degner SC, Verma-Gaur J, Wong TP, Bossen C, Iverson GM, Torkamani A, Vettermann C, Lin YC, Ju Z, Schulz D, et al. CCCTC-binding factor (CTCF) and cohesin influence the genomic architecture of the Igh locus and antisense transcription in pro-B cells. Proc. Natl. Acad. Sci. USA. 2011;108:9566–9571. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

