Recurrent HERV-H-mediated 3q13.2-q13.31 deletions cause a syndrome of hypotonia and motor, language, and cognitive delays
- PMID: 23878096
- PMCID: PMC4599348
- DOI: 10.1002/humu.22384
Recurrent HERV-H-mediated 3q13.2-q13.31 deletions cause a syndrome of hypotonia and motor, language, and cognitive delays
Abstract
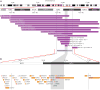
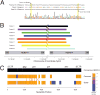
We describe the molecular and clinical characterization of nine individuals with recurrent, 3.4-Mb, de novo deletions of 3q13.2-q13.31 detected by chromosomal microarray analysis. All individuals have hypotonia and language and motor delays; they variably express mild to moderate cognitive delays (8/9), abnormal behavior (7/9), and autism spectrum disorders (3/9). Common facial features include downslanting palpebral fissures with epicanthal folds, a slightly bulbous nose, and relative macrocephaly. Twenty-eight genes map to the deleted region, including four strong candidate genes, DRD3, ZBTB20, GAP43, and BOC, with important roles in neural and/or muscular development. Analysis of the breakpoint regions based on array data revealed directly oriented human endogenous retrovirus (HERV-H) elements of ~5 kb in size and of >95% DNA sequence identity flanking the deletion. Subsequent DNA sequencing revealed different deletion breakpoints and suggested nonallelic homologous recombination (NAHR) between HERV-H elements as a mechanism of deletion formation, analogous to HERV-I-flanked and NAHR-mediated AZFa deletions. We propose that similar HERV elements may also mediate other recurrent deletion and duplication events on a genome-wide scale. Observation of rare recurrent chromosomal events such as these deletions helps to further the understanding of mechanisms behind naturally occurring variation in the human genome and its contribution to genetic disease.
Keywords: 3q13; HERV-H; NAHR; developmental delay; hypotonia; microarray; microdeletion; recurrent.
© 2013 WILEY PERIODICALS, INC.
Figures

Similar articles
-
Female patient with autistic disorder, intellectual disability, and co-morbid anxiety disorder: Expanding the phenotype associated with the recurrent 3q13.2-q13.31 microdeletion.Am J Med Genet A. 2015 Dec;167A(12):3121-9. doi: 10.1002/ajmg.a.37292. Epub 2015 Aug 29. Am J Med Genet A. 2015. PMID: 26332054
-
Human endogenous retroviral elements promote genome instability via non-allelic homologous recombination.BMC Biol. 2014 Sep 23;12:74. doi: 10.1186/s12915-014-0074-4. BMC Biol. 2014. PMID: 25246103 Free PMC article.
-
Expanding the clinical phenotype at the 3q13.31 locus with a new case of microdeletion and first characterization of the reciprocal duplication.Mol Genet Metab. 2013 Sep-Oct;110(1-2):90-7. doi: 10.1016/j.ymgme.2013.07.013. Epub 2013 Jul 20. Mol Genet Metab. 2013. PMID: 23920044
-
Chromosome 15q24 microdeletion syndrome.Orphanet J Rare Dis. 2012 Jan 4;7:2. doi: 10.1186/1750-1172-7-2. Orphanet J Rare Dis. 2012. PMID: 22216833 Free PMC article. Review.
-
1q21.3 deletion involving GATAD2B: An emerging recurrent microdeletion syndrome.Am J Med Genet A. 2017 Mar;173(3):766-770. doi: 10.1002/ajmg.a.38082. Am J Med Genet A. 2017. PMID: 28211977 Review.
Cited by
-
Developmental and adult GAP-43 deficiency in mice dynamically alters hippocampal neurogenesis and mossy fiber volume.Dev Neurosci. 2014;36(1):44-63. doi: 10.1159/000357840. Epub 2014 Feb 26. Dev Neurosci. 2014. PMID: 24576816 Free PMC article.
-
Boc modifies the spectrum of holoprosencephaly in the absence of Gas1 function.Biol Open. 2014 Jul 25;3(8):728-40. doi: 10.1242/bio.20147989. Biol Open. 2014. PMID: 25063195 Free PMC article.
-
Recurrent 8q13.2-13.3 microdeletions associated with branchio-oto-renal syndrome are mediated by human endogenous retroviral (HERV) sequence blocks.BMC Med Genet. 2014 Aug 19;15:90. doi: 10.1186/s12881-014-0090-9. BMC Med Genet. 2014. PMID: 25135225 Free PMC article.
-
Unbalanced translocations arise from diverse mutational mechanisms including chromothripsis.Genome Res. 2015 Jul;25(7):937-47. doi: 10.1101/gr.191247.115. Epub 2015 Jun 12. Genome Res. 2015. PMID: 26070663 Free PMC article.
-
Genome architecture and its roles in human copy number variation.Genomics Inform. 2014 Dec;12(4):136-44. doi: 10.5808/GI.2014.12.4.136. Epub 2014 Dec 31. Genomics Inform. 2014. PMID: 25705150 Free PMC article. Review.
References
-
- Arai K, Matukiyo H, Takazawa H. A case report of partial deletion of the long arm of the no 3 chromosome. Med Genet Res. 1982;4:1–4.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

