Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations
- PMID: 22820256
- PMCID: PMC3413789
- DOI: 10.1038/nature11329
Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations
Abstract
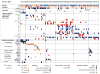
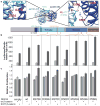
Medulloblastomas are the most common malignant brain tumours in children. Identifying and understanding the genetic events that drive these tumours is critical for the development of more effective diagnostic, prognostic and therapeutic strategies. Recently, our group and others described distinct molecular subtypes of medulloblastoma on the basis of transcriptional and copy number profiles. Here we use whole-exome hybrid capture and deep sequencing to identify somatic mutations across the coding regions of 92 primary medulloblastoma/normal pairs. Overall, medulloblastomas have low mutation rates consistent with other paediatric tumours, with a median of 0.35 non-silent mutations per megabase. We identified twelve genes mutated at statistically significant frequencies, including previously known mutated genes in medulloblastoma such as CTNNB1, PTCH1, MLL2, SMARCA4 and TP53. Recurrent somatic mutations were newly identified in an RNA helicase gene, DDX3X, often concurrent with CTNNB1 mutations, and in the nuclear co-repressor (N-CoR) complex genes GPS2, BCOR and LDB1. We show that mutant DDX3X potentiates transactivation of a TCF promoter and enhances cell viability in combination with mutant, but not wild-type, β-catenin. Together, our study reveals the alteration of WNT, hedgehog, histone methyltransferase and now N-CoR pathways across medulloblastomas and within specific subtypes of this disease, and nominates the RNA helicase DDX3X as a component of pathogenic β-catenin signalling in medulloblastoma.
Figures

Similar articles
-
Dissecting the genomic complexity underlying medulloblastoma.Nature. 2012 Aug 2;488(7409):100-5. doi: 10.1038/nature11284. Nature. 2012. PMID: 22832583 Free PMC article.
-
Novel mutations target distinct subgroups of medulloblastoma.Nature. 2012 Aug 2;488(7409):43-8. doi: 10.1038/nature11213. Nature. 2012. PMID: 22722829 Free PMC article.
-
Medulloblastoma in the age of molecular subgroups: a review.J Neurosurg Pediatr. 2019 Oct 1;24(4):353-363. doi: 10.3171/2019.5.PEDS18381. Epub 2019 Oct 1. J Neurosurg Pediatr. 2019. PMID: 31574483 Review.
-
Genome sequencing of SHH medulloblastoma predicts genotype-related response to smoothened inhibition.Cancer Cell. 2014 Mar 17;25(3):393-405. doi: 10.1016/j.ccr.2014.02.004. Cancer Cell. 2014. PMID: 24651015 Free PMC article.
-
The Role of the RNA Helicase DDX3X in Medulloblastoma Progression.Biomolecules. 2024 Jul 6;14(7):803. doi: 10.3390/biom14070803. Biomolecules. 2024. PMID: 39062517 Free PMC article. Review.
Cited by
-
Aberrant patterns of H3K4 and H3K27 histone lysine methylation occur across subgroups in medulloblastoma.Acta Neuropathol. 2013 Mar;125(3):373-84. doi: 10.1007/s00401-012-1070-9. Epub 2012 Nov 25. Acta Neuropathol. 2013. PMID: 23184418 Free PMC article.
-
Molecular Classification of Medulloblastoma.Neurol Med Chir (Tokyo). 2016 Nov 15;56(11):687-697. doi: 10.2176/nmc.ra.2016-0016. Epub 2016 May 26. Neurol Med Chir (Tokyo). 2016. PMID: 27238212 Free PMC article. Review.
-
Medulloblastoma biology in the post-genomic era.Future Oncol. 2012 Dec;8(12):1597-604. doi: 10.2217/fon.12.151. Future Oncol. 2012. PMID: 23231521 Free PMC article. Review.
-
Medulloblastoma development: tumor biology informs treatment decisions.CNS Oncol. 2015;4(2):79-89. doi: 10.2217/cns.14.58. CNS Oncol. 2015. PMID: 25768332 Free PMC article. Review.
-
Whole genome and transcriptome sequencing of a B3 thymoma.PLoS One. 2013;8(4):e60572. doi: 10.1371/journal.pone.0060572. Epub 2013 Apr 5. PLoS One. 2013. PMID: 23577124 Free PMC article.
References
-
- CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2004–2007. Central Brain Tumor Registry of the United States; Hinsdale, IL: 2011. at < www.cbtrus.org>.
-
- Remke M, et al. Adult medulloblastoma comprises three major molecular variants. J Clin Oncol. 2011;29:2717–2723. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- L40 NS063706/NS/NINDS NIH HHS/United States
- U54 HG003067/HG/NHGRI NIH HHS/United States
- R01 NS046789/NS/NINDS NIH HHS/United States
- CAPMC/ CIHR/Canada
- HHMI/Howard Hughes Medical Institute/United States
- U54HG003067/HG/NHGRI NIH HHS/United States
- R01 CA105607/CA/NCI NIH HHS/United States
- P30 HD018655/HD/NICHD NIH HHS/United States
- R01CA105607/CA/NCI NIH HHS/United States
- R01 CA030002/CA/NCI NIH HHS/United States
- P01 CA050661/CA/NCI NIH HHS/United States
- R25NS070682/NS/NINDS NIH HHS/United States
- R01 CA109467/CA/NCI NIH HHS/United States
- R01 CA148699/CA/NCI NIH HHS/United States
- R01CA148699/CA/NCI NIH HHS/United States
- CA050661/CA/NCI NIH HHS/United States
- R01 CA154480/CA/NCI NIH HHS/United States
- R25 NS070682/NS/NINDS NIH HHS/United States
- R01CA109467/CA/NCI NIH HHS/United States
- P30 HD18655/HD/NICHD NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

