Evolutionarily conserved substrate substructures for automated annotation of enzyme superfamilies
- PMID: 18670595
- PMCID: PMC2453236
- DOI: 10.1371/journal.pcbi.1000142
Evolutionarily conserved substrate substructures for automated annotation of enzyme superfamilies
Abstract
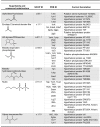
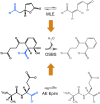
The evolution of enzymes affects how well a species can adapt to new environmental conditions. During enzyme evolution, certain aspects of molecular function are conserved while other aspects can vary. Aspects of function that are more difficult to change or that need to be reused in multiple contexts are often conserved, while those that vary may indicate functions that are more easily changed or that are no longer required. In analogy to the study of conservation patterns in enzyme sequences and structures, we have examined the patterns of conservation and variation in enzyme function by analyzing graph isomorphisms among enzyme substrates of a large number of enzyme superfamilies. This systematic analysis of substrate substructures establishes the conservation patterns that typify individual superfamilies. Specifically, we determined the chemical substructures that are conserved among all known substrates of a superfamily and the substructures that are reacting in these substrates and then examined the relationship between the two. Across the 42 superfamilies that were analyzed, substantial variation was found in how much of the conserved substructure is reacting, suggesting that superfamilies may not be easily grouped into discrete and separable categories. Instead, our results suggest that many superfamilies may need to be treated individually for analyses of evolution, function prediction, and guiding enzyme engineering strategies. Annotating superfamilies with these conserved and reacting substructure patterns provides information that is orthogonal to information provided by studies of conservation in superfamily sequences and structures, thereby improving the precision with which we can predict the functions of enzymes of unknown function and direct studies in enzyme engineering. Because the method is automated, it is suitable for large-scale characterization and comparison of fundamental functional capabilities of both characterized and uncharacterized enzyme superfamilies.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
A strategy for large-scale comparison of evolutionary- and reaction-based classifications of enzyme function.Database (Oxford). 2020 Jan 1;2020:baaa034. doi: 10.1093/database/baaa034. Database (Oxford). 2020. PMID: 32449511 Free PMC article.
-
Evolution of function in protein superfamilies, from a structural perspective.J Mol Biol. 2001 Apr 6;307(4):1113-43. doi: 10.1006/jmbi.2001.4513. J Mol Biol. 2001. PMID: 11286560
-
A gold standard set of mechanistically diverse enzyme superfamilies.Genome Biol. 2006;7(1):R8. doi: 10.1186/gb-2006-7-1-r8. Epub 2006 Jan 31. Genome Biol. 2006. PMID: 16507141 Free PMC article.
-
Divergence and convergence in enzyme evolution.J Biol Chem. 2012 Jan 2;287(1):21-28. doi: 10.1074/jbc.R111.241976. Epub 2011 Nov 8. J Biol Chem. 2012. PMID: 22069324 Free PMC article. Review.
-
Evolution of Enzyme Superfamilies: Comprehensive Exploration of Sequence-Function Relationships.Biochemistry. 2016 Nov 22;55(46):6375-6388. doi: 10.1021/acs.biochem.6b00723. Epub 2016 Nov 11. Biochemistry. 2016. PMID: 27802036 Review.
Cited by
-
Quantitative comparison of catalytic mechanisms and overall reactions in convergently evolved enzymes: implications for classification of enzyme function.PLoS Comput Biol. 2010 Mar 12;6(3):e1000700. doi: 10.1371/journal.pcbi.1000700. PLoS Comput Biol. 2010. PMID: 20300652 Free PMC article.
-
A strategy for large-scale comparison of evolutionary- and reaction-based classifications of enzyme function.Database (Oxford). 2020 Jan 1;2020:baaa034. doi: 10.1093/database/baaa034. Database (Oxford). 2020. PMID: 32449511 Free PMC article.
-
Accuracy of functional surfaces on comparatively modeled protein structures.J Struct Funct Genomics. 2011 Jul;12(2):97-107. doi: 10.1007/s10969-011-9109-z. Epub 2011 May 4. J Struct Funct Genomics. 2011. PMID: 21541664 Free PMC article.
-
Characterizing the complexity of enzymes on the basis of their mechanisms and structures with a bio-computational analysis.FEBS J. 2011 Oct;278(20):3835-45. doi: 10.1111/j.1742-4658.2011.08190.x. Epub 2011 Jun 13. FEBS J. 2011. PMID: 21605342 Free PMC article. Review.
-
FINDSITE: a threading-based approach to ligand homology modeling.PLoS Comput Biol. 2009 Jun;5(6):e1000405. doi: 10.1371/journal.pcbi.1000405. Epub 2009 Jun 5. PLoS Comput Biol. 2009. PMID: 19503616 Free PMC article.
References
-
- Babbitt PC, Gerlt JA. Understanding enzyme superfamilies. Chemistry as the fundamental determinant in the evolution of new catalytic activities. J Biol Chem. 1997;272:30591–30594. - PubMed
-
- Copley SD. Evolution of a metabolic pathway for degradation of a toxic xenobiotic: the patchwork approach. Trends Biochem Sci. 2000;25:261–265. - PubMed
-
- Aharoni A, Gaidukov L, Khersonsky O, McQ Gould S, Roodveldt C, et al. The ‘evolvability’ of promiscuous protein functions. Nat Genet. 2005;37:73–76. - PubMed
-
- Riesenfeld CS, Schloss PD, Handelsman J. Metagenomics: genomic analysis of microbial communities. Annu Rev Genet. 2004;38:525–552. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

