A novel function for the N-terminal nucleophile hydrolase fold demonstrated by the structure of an archaeal inosine monophosphate cyclohydrolase
- PMID: 17407260
- PMCID: PMC2631436
- DOI: 10.1021/bi061637j
A novel function for the N-terminal nucleophile hydrolase fold demonstrated by the structure of an archaeal inosine monophosphate cyclohydrolase
Abstract
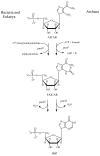
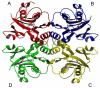
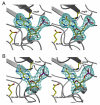
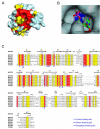
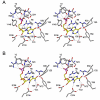
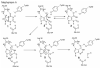
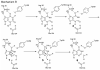
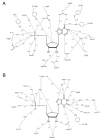
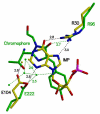
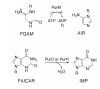
Inosine 5'-monophosphate (IMP) cyclohydrolase catalyzes the cyclization of 5-formaminoimidazole-4-carboxamide ribonucleotide (FAICAR) to IMP in the final step of de novo purine biosynthesis. Two major types of this enzyme have been discovered to date: PurH in Bacteria and Eukarya and PurO in Archaea. The structure of the MTH1020 gene product from Methanothermobacter thermoautotrophicus was previously solved without functional annotation but shows high amino acid sequence similarity to other PurOs. We determined the crystal structure of the MTH1020 gene product in complex with either IMP or 5-aminoimidazole-4-carboxamide ribonucleotide (AICAR) at 2.0 and 2.6 A resolution, respectively. On the basis of the sequence analysis, ligand-bound structures, and biochemical data, MTH1020 is confirmed as an archaeal IMP cyclohydrolase, thus designated as MthPurO. MthPurO has a four-layered alphabeta betaalpha core structure, showing an N-terminal nucleophile (NTN) hydrolase fold. The active site is located at the deep pocket between two central beta-sheets and contains residues strictly conserved within PurOs. Comparisons of the two types of IMP cyclohydrolase, PurO and PurH, revealed that there are no similarities in sequence, structure, or the active site architecture, suggesting that they are evolutionarily not related to each other. The MjR31K mutant of PurO from Methanocaldococcus jannaschii showed 76% decreased activity and the MjE102Q mutation completely abolished enzymatic activity, suggesting that these highly conserved residues play critical roles in catalysis. Interestingly, green fluorescent protein (GFP), which has no structural homology to either PurO or PurH but catalyzes a similar intramolecular cyclohydrolase reaction required for chromophore maturation, utilizes Arg96 and Glu222 in a mechanism analogous to that of PurO.
Figures
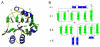
Similar articles
-
Structural analyses of a purine biosynthetic enzyme from Mycobacterium tuberculosis reveal a novel bound nucleotide.J Biol Chem. 2011 Nov 25;286(47):40706-16. doi: 10.1074/jbc.M111.291138. Epub 2011 Sep 28. J Biol Chem. 2011. PMID: 21956117 Free PMC article.
-
Characterization of AICAR transformylase/IMP cyclohydrolase (ATIC) from Staphylococcus lugdunensis.FEBS J. 2017 Dec;284(24):4233-4261. doi: 10.1111/febs.14303. Epub 2017 Nov 13. FEBS J. 2017. PMID: 29063699
-
Catalytic mechanism of the cyclohydrolase activity of human aminoimidazole carboxamide ribonucleotide formyltransferase/inosine monophosphate cyclohydrolase.Biochemistry. 2004 Feb 10;43(5):1184-92. doi: 10.1021/bi035139b. Biochemistry. 2004. PMID: 14756554
-
Modulation of the de novo purine nucleotide pathway as a therapeutic strategy in mitochondrial myopathy.Pharmacol Res. 2018 Dec;138:37-42. doi: 10.1016/j.phrs.2018.09.027. Epub 2018 Sep 26. Pharmacol Res. 2018. PMID: 30267763 Review.
-
[The second half of de novo synthetic pathway of IMP and conversion of IMP to AMP].Nihon Rinsho. 2003 Jan;61 Suppl 1:52-8. Nihon Rinsho. 2003. PMID: 12629690 Review. Japanese. No abstract available.
Cited by
-
Purine biosynthesis in archaea: variations on a theme.Biol Direct. 2011 Dec 14;6:63. doi: 10.1186/1745-6150-6-63. Biol Direct. 2011. PMID: 22168471 Free PMC article.
-
Different Ways of Doing the Same: Variations in the Two Last Steps of the Purine Biosynthetic Pathway in Prokaryotes.Genome Biol Evol. 2019 Apr 1;11(4):1235-1249. doi: 10.1093/gbe/evz035. Genome Biol Evol. 2019. PMID: 30785193 Free PMC article.
-
Structural analyses of a purine biosynthetic enzyme from Mycobacterium tuberculosis reveal a novel bound nucleotide.J Biol Chem. 2011 Nov 25;286(47):40706-16. doi: 10.1074/jbc.M111.291138. Epub 2011 Sep 28. J Biol Chem. 2011. PMID: 21956117 Free PMC article.
-
Comparative genomics of nucleotide metabolism: a tour to the past of the three cellular domains of life.BMC Genomics. 2014 Sep 17;15(1):800. doi: 10.1186/1471-2164-15-800. BMC Genomics. 2014. PMID: 25230797 Free PMC article.
-
Initial insight into the function of the lysosomal 66.3 kDa protein from mouse by means of X-ray crystallography.BMC Struct Biol. 2009 Aug 25;9:56. doi: 10.1186/1472-6807-9-56. BMC Struct Biol. 2009. PMID: 19706171 Free PMC article.
References
-
- Rayl EA, Moroson BA, Beardsley GP. The human purH gene product, 5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase. Cloning, sequencing, expression, purification, kinetic analysis, and domain mapping. J. Biol. Chem. 1996;271:2225–2233. - PubMed
-
- Greasley SE, Horton P, Ramcharan J, Beardsley GP, Benkovic SJ, Wilson IA. Crystal structure of a bifunctional transformylase and cyclohydrolase enzyme in purine biosynthesis. Nat. Struct. Biol. 2001;8:402–406. - PubMed
-
- Wolan DW, Cheong CG, Greasley SE, Wilson IA. Structural insights into the human and avian IMP cyclohydrolase mechanism via crystal structures with the bound XMP inhibitor. Biochemistry. 2004;43:1171–1183. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

