Evidence for multiple levels of regulation of Oenococcus oeni clpP-clpL locus expression in response to stress
- PMID: 15028706
- PMCID: PMC374425
- DOI: 10.1128/JB.186.7.2200-2205.2003
Evidence for multiple levels of regulation of Oenococcus oeni clpP-clpL locus expression in response to stress
Abstract
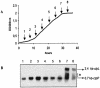
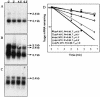
A locus containing the clpP and clpL genes in the lactic acid bacterium Oenococcus oeni was studied. Real-time reverse transcription-PCR analysis revealed different induction factors involved in expression of these genes during stress. According to the conditions, clpP and clpL genes could be transcripted as two distinct transcripts or cotranscripted. The clpP promoter depended on the CtsR regulator, but surprisingly the clpL promoter did not. The amount of the clpL transcript depended on mRNA stability. This clp ATPase gene is at least controlled at the posttranscriptional level.
Figures

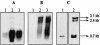

Similar articles
-
The Oenococcus oeni clpX homologue is a heat shock gene preferentially expressed in exponential growth phase.J Bacteriol. 1999 Nov;181(21):6634-41. doi: 10.1128/JB.181.21.6634-6641.1999. J Bacteriol. 1999. PMID: 10542163 Free PMC article.
-
Stress induction of the Bacillus subtilis clpP gene encoding a homologue of the proteolytic component of the Clp protease and the involvement of ClpP and ClpX in stress tolerance.Mol Microbiol. 1998 May;28(4):787-802. doi: 10.1046/j.1365-2958.1998.00840.x. Mol Microbiol. 1998. PMID: 9643546
-
CtsR is the master regulator of stress response gene expression in Oenococcus oeni.J Bacteriol. 2005 Aug;187(16):5614-23. doi: 10.1128/JB.187.16.5614-5623.2005. J Bacteriol. 2005. PMID: 16077106 Free PMC article.
-
Regulation of stress response in Oenococcus oeni as a function of environmental changes and growth phase.Int J Food Microbiol. 2000 Apr 10;55(1-3):27-31. doi: 10.1016/s0168-1605(00)00209-9. Int J Food Microbiol. 2000. PMID: 10791713 Review.
-
A regulatory switch involving a Clp ATPase.Bioessays. 1997 Jun;19(6):455-8. doi: 10.1002/bies.950190604. Bioessays. 1997. PMID: 9204762 Review.
Cited by
-
The early response to acid shock in Lactobacillus reuteri involves the ClpL chaperone and a putative cell wall-altering esterase.Appl Environ Microbiol. 2007 Jun;73(12):3924-35. doi: 10.1128/AEM.01502-06. Epub 2007 Apr 20. Appl Environ Microbiol. 2007. PMID: 17449683 Free PMC article.
-
Significant influence of four highly conserved amino-acids in lipochaperon-active sHsps on the structure and functions of the Lo18 protein.Sci Rep. 2023 Nov 3;13(1):19036. doi: 10.1038/s41598-023-46306-6. Sci Rep. 2023. PMID: 37923897 Free PMC article.
-
Transcriptomic and Proteomic Analysis of Oenococcus oeni Adaptation to Wine Stress Conditions.Front Microbiol. 2016 Sep 30;7:1554. doi: 10.3389/fmicb.2016.01554. eCollection 2016. Front Microbiol. 2016. PMID: 27746771 Free PMC article.
-
CtsR regulation in mcsAB-deficient Gram-positive bacteria.J Bacteriol. 2012 Mar;194(6):1361-8. doi: 10.1128/JB.06746-11. Epub 2012 Jan 13. J Bacteriol. 2012. PMID: 22247503 Free PMC article.
-
Selection and Validation of Reference Genes for Quantitative Real-Time PCR Normalization Under Ethanol Stress Conditions in Oenococcus oeni SD-2a.Front Microbiol. 2018 May 4;9:892. doi: 10.3389/fmicb.2018.00892. eCollection 2018. Front Microbiol. 2018. PMID: 29780378 Free PMC article.
References
-
- Cavin, J. F., P. Schmitt, A. Arias, J. Lin, and C. Divies. 1988. Plasmid profiles in Leuconostoc species. Microbiol. Aliment. Nutr. 5:55-62.
-
- Derre, I., G. Rapoport, K. Devine, M. Rose, and T. Msadek. 1999. ClpE, a novel type of HSP100 ATPase, is part of the CtsR heat shock regulon of Bacillus subtilis. Mol. Microbiol. 32:581-593. - PubMed
-
- Derre, I., G. Rapoport, and T. Msadek. 1999. CtsR, a novel regulator of stress and heat shock response, controls clp and molecular chaperone gene expression in Gram-positive bacteria. Mol. Microbiol. 31:117-131. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Miscellaneous

