Duane radial ray syndrome (Okihiro syndrome) maps to 20q13 and results from mutations in SALL4, a new member of the SAL family
- PMID: 12395297
- PMCID: PMC385096
- DOI: 10.1086/343821
Duane radial ray syndrome (Okihiro syndrome) maps to 20q13 and results from mutations in SALL4, a new member of the SAL family
Abstract
Duane syndrome is a congenital eye movement disorder characterized most typically by absence of abduction, restricted adduction, and retraction of the globe on attempted adduction. Duane syndrome can be coinherited with radial ray anomalies as an autosomal dominant trait, referred to as "Okihiro syndrome" or "Duane radial ray syndrome" (DRRS). We ascertained three pedigrees with DRRS and mapped their disease gene to a 21.6-cM region of chromosome 20 flanked by markers D20S888 and D20S102. A new member of the SAL family of proposed C(2)H(2) zinc finger transcription factors, SALL4, falls within the region. Mutation analysis of SALL4 in the three pedigrees revealed one nonsense and two frameshift heterozygous mutations. SALL4 represents the first identified Duane syndrome gene and the second malformation syndrome resulting from mutations in SAL genes and likely plays a critical role in abducens motoneuron development.
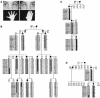
Figures

Similar articles
-
SALL4 mutations in Okihiro syndrome (Duane-radial ray syndrome), acro-renal-ocular syndrome, and related disorders.Hum Mutat. 2005 Sep;26(3):176-83. doi: 10.1002/humu.20215. Hum Mutat. 2005. PMID: 16086360
-
Okihiro syndrome is caused by SALL4 mutations.Hum Mol Genet. 2002 Nov 1;11(23):2979-87. doi: 10.1093/hmg/11.23.2979. Hum Mol Genet. 2002. PMID: 12393809
-
Novel frameshift variant in gene SALL4 causing Okihiro syndrome.Eur J Med Genet. 2016 Feb;59(2):80-5. doi: 10.1016/j.ejmg.2015.12.015. Epub 2016 Jan 11. Eur J Med Genet. 2016. PMID: 26791099
-
[Progress of study on the transcription factor SALL4].Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2011 Jun;19(3):820-3. Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2011. PMID: 21729579 Review. Chinese.
-
HOXA1 mutations are not a common cause of Duane anomaly.Am J Med Genet A. 2006 Apr 15;140(8):900-2. doi: 10.1002/ajmg.a.31167. Am J Med Genet A. 2006. PMID: 16528738 Free PMC article. Review. No abstract available.
Cited by
-
SALL4 is a CRL3REN/KCTD11 substrate that drives Sonic Hedgehog-dependent medulloblastoma.Cell Death Differ. 2024 Feb;31(2):170-187. doi: 10.1038/s41418-023-01246-6. Epub 2023 Dec 7. Cell Death Differ. 2024. PMID: 38062245 Free PMC article.
-
Clinical geneticists' views of VACTERL/VATER association.Am J Med Genet A. 2012 Dec;158A(12):3087-100. doi: 10.1002/ajmg.a.35638. Epub 2012 Nov 19. Am J Med Genet A. 2012. PMID: 23165726 Free PMC article. Review.
-
Congenital abnormalities of cranial nerve development: overview, molecular mechanisms, and further evidence of heterogeneity and complexity of syndromes with congenital limitation of eye movements.Trans Am Ophthalmol Soc. 2004;102:373-89. Trans Am Ophthalmol Soc. 2004. PMID: 15747768 Free PMC article.
-
Magnetic resonance imaging of innervational and extraocular muscle abnormalities in Duane-radial ray syndrome.Invest Ophthalmol Vis Sci. 2007 Dec;48(12):5505-11. doi: 10.1167/iovs.07-0772. Invest Ophthalmol Vis Sci. 2007. PMID: 18055799 Free PMC article.
-
Differential roles of Sall4 isoforms in embryonic stem cell pluripotency.Mol Cell Biol. 2010 Nov;30(22):5364-80. doi: 10.1128/MCB.00419-10. Epub 2010 Sep 13. Mol Cell Biol. 2010. PMID: 20837710 Free PMC article.
References
Electronic-Database Information
-
- ExPASy Molecular Biology Server, http://us.expasy.org/ (for prediction programs SWISS-PROT and TrEMBL)
-
- GenBank, http://www.ncbi.nlm.nih.gov/Genbank/ (for SALL4 genomic, NT_011362; SALL4 cDNA, NM_020436; SALL4, NP_065169; SALL1, NP_002959; SALL2, XP_033473; SALL3, AAK18311; Xsal-3, BAA85900)
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for DURS1, including DRRS, [MIM 126800] and DURS2 [MIM 604356])
-
- UCSC Human Genome Project Working Draft, http://genome.cse.ucsc.edu/ (for working drafts for the mouse genome and the human genome)
References
-
- Appukuttan B, Gillanders E, Juo SH, Freas-Lutz D, Ott S, Sood R, Van Auken A, Bailey-Wilson J, Wang X, Patel RJ, Robbins CM, Chung M, Annett G, Weinberg K, Borchert MS, Trent JM, Brownstein MJ, Stout JT (1999) Localization of a gene for Duane retraction syndrome to chromosome 2q31. Am J Hum Genet 65:1639–1646 - PMC - PubMed
-
- Chun BB, Mazzoli RA, Raymond WR (2001) Characteristics of Okihiro syndrome. J Pediatr Ophthalmol Strabismus 38:235–239 - PubMed
-
- Chung M, Stout JT, Borchert MS (2000) Clinical diversity of hereditary Duane's retraction syndrome. Ophthalmology 107:500–503 - PubMed
-
- Engle EC, Kunkel LM, Specht LA, Beggs AH (1994) Mapping a gene for congenital fibrosis of the extraocular muscles to the centromeric region of chromosome 12. Nat Genet 7:69–73 - PubMed
-
- Hotchkiss MG, Miller NR, Clark AW, Green WG (1980) Bilateral Duane's retraction syndrome: a clinical-pathological case report. Arch Ophthalmol 98:870–874 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

