A maternally methylated CpG island in KvLQT1 is associated with an antisense paternal transcript and loss of imprinting in Beckwith-Wiedemann syndrome
- PMID: 10393948
- PMCID: PMC22188
- DOI: 10.1073/pnas.96.14.8064
A maternally methylated CpG island in KvLQT1 is associated with an antisense paternal transcript and loss of imprinting in Beckwith-Wiedemann syndrome
Abstract
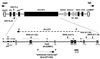
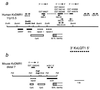
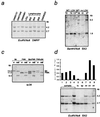
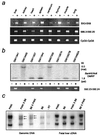
Loss of imprinting at IGF2, generally through an H19-independent mechanism, is associated with a large percentage of patients with the overgrowth and cancer predisposition condition Beckwith-Wiedemann syndrome (BWS). Imprinting control elements are proposed to exist within the KvLQT1 locus, because multiple BWS-associated chromosome rearrangements disrupt this gene. We have identified an evolutionarily conserved, maternally methylated CpG island (KvDMR1) in an intron of the KvLQT1 gene. Among 12 cases of BWS with normal H19 methylation, 5 showed demethylation of KvDMR1 in fibroblast or lymphocyte DNA; whereas, in 4 cases of BWS with H19 hypermethylation, methylation at KvDMRl was normal. Thus, inactivation of H19 and hypomethylation at KvDMR1 (or an associated phenomenon) represent distinct epigenetic anomalies associated with biallelic expression of IGF2. Reverse transcription-PCR analysis of the human and syntenic mouse loci identified the presence of a KvDMR1-associated RNA transcribed exclusively from the paternal allele and in the opposite orientation with respect to the maternally expressed KvLQT1 gene. We propose that KvDMR1 and/or its associated antisense RNA (KvLQT1-AS) represents an additional imprinting control element or center in the human 11p15.5 and mouse distal 7 imprinted domains.
Figures
Similar articles
-
Relaxation of insulin-like growth factor 2 imprinting and discordant methylation at KvDMR1 in two first cousins affected by Beckwith-Wiedemann and Klippel-Trenaunay-Weber syndromes.Am J Hum Genet. 2000 Mar;66(3):841-7. doi: 10.1086/302811. Am J Hum Genet. 2000. PMID: 10712200 Free PMC article.
-
Loss of imprinting of a paternally expressed transcript, with antisense orientation to KVLQT1, occurs frequently in Beckwith-Wiedemann syndrome and is independent of insulin-like growth factor II imprinting.Proc Natl Acad Sci U S A. 1999 Apr 27;96(9):5203-8. doi: 10.1073/pnas.96.9.5203. Proc Natl Acad Sci U S A. 1999. PMID: 10220444 Free PMC article.
-
Analysis of the methylation status of the KCNQ1OT and H19 genes in leukocyte DNA for the diagnosis and prognosis of Beckwith-Wiedemann syndrome.Eur J Hum Genet. 2001 Jun;9(6):409-18. doi: 10.1038/sj.ejhg.5200649. Eur J Hum Genet. 2001. PMID: 11436121
-
Epigenetic and genetic alterations of the imprinting disorder Beckwith-Wiedemann syndrome and related disorders.J Hum Genet. 2013 Jul;58(7):402-9. doi: 10.1038/jhg.2013.51. Epub 2013 May 30. J Hum Genet. 2013. PMID: 23719190 Review.
-
Beckwith-Wiedemann syndrome.Am J Med Genet C Semin Med Genet. 2010 Aug 15;154C(3):343-54. doi: 10.1002/ajmg.c.30267. Am J Med Genet C Semin Med Genet. 2010. PMID: 20803657 Review.
Cited by
-
The role of CTCF in the organization of the centromeric 11p15 imprinted domain interactome.Nucleic Acids Res. 2021 Jun 21;49(11):6315-6330. doi: 10.1093/nar/gkab475. Nucleic Acids Res. 2021. PMID: 34107024 Free PMC article.
-
G9a histone methyltransferase contributes to imprinting in the mouse placenta.Mol Cell Biol. 2008 Feb;28(3):1104-13. doi: 10.1128/MCB.01111-07. Epub 2007 Nov 26. Mol Cell Biol. 2008. PMID: 18039842 Free PMC article.
-
Relaxation of insulin-like growth factor 2 imprinting and discordant methylation at KvDMR1 in two first cousins affected by Beckwith-Wiedemann and Klippel-Trenaunay-Weber syndromes.Am J Hum Genet. 2000 Mar;66(3):841-7. doi: 10.1086/302811. Am J Hum Genet. 2000. PMID: 10712200 Free PMC article.
-
Induction of p57(KIP2) expression by p73beta.Proc Natl Acad Sci U S A. 2002 Mar 19;99(6):3529-34. doi: 10.1073/pnas.062491899. Epub 2002 Mar 12. Proc Natl Acad Sci U S A. 2002. PMID: 11891335 Free PMC article.
-
Fate of induced pluripotent stem cells following transplantation to murine seminiferous tubules.Hum Mol Genet. 2014 Jun 15;23(12):3071-84. doi: 10.1093/hmg/ddu012. Epub 2014 Jan 20. Hum Mol Genet. 2014. PMID: 24449759 Free PMC article.
References
-
- Barlow D P. Trends Genet. 1994;10:194–199. - PubMed
-
- Constancia M, Pickard B, Kelsey G, Reik W. Genome Res. 1998;8:881–900. - PubMed
-
- Surani M A. Cell. 1998;93:309–312. - PubMed
-
- Li E, Beard C, Jaenisch R. Nature (London) 1993;366:362–365. - PubMed
-
- Olek A, Walter J. Nat Genet. 1997;17:275–276. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

